IL-10 constrains sphingolipid metabolism to limit inflammation
- PMID: 38383790
- PMCID: PMC10954550
- DOI: 10.1038/s41586-024-07098-5
IL-10 constrains sphingolipid metabolism to limit inflammation
Abstract
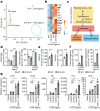
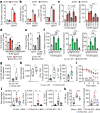
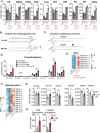
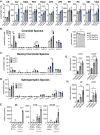
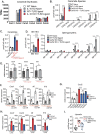
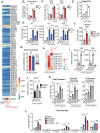
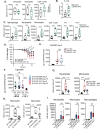
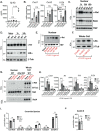
Interleukin-10 (IL-10) is a key anti-inflammatory cytokine that can limit immune cell activation and cytokine production in innate immune cell types1. Loss of IL-10 signalling results in life-threatening inflammatory bowel disease in humans and mice-however, the exact mechanism by which IL-10 signalling subdues inflammation remains unclear2-5. Here we find that increased saturated very long chain (VLC) ceramides are critical for the heightened inflammatory gene expression that is a hallmark of IL-10 deficiency. Accordingly, genetic deletion of ceramide synthase 2 (encoded by Cers2), the enzyme responsible for VLC ceramide production, limited the exacerbated inflammatory gene expression programme associated with IL-10 deficiency both in vitro and in vivo. The accumulation of saturated VLC ceramides was regulated by a decrease in metabolic flux through the de novo mono-unsaturated fatty acid synthesis pathway. Restoring mono-unsaturated fatty acid availability to cells deficient in IL-10 signalling limited saturated VLC ceramide production and the associated inflammation. Mechanistically, we find that persistent inflammation mediated by VLC ceramides is largely dependent on sustained activity of REL, an immuno-modulatory transcription factor. Together, these data indicate that an IL-10-driven fatty acid desaturation programme rewires VLC ceramide accumulation and aberrant activation of REL. These studies support the idea that fatty acid homeostasis in innate immune cells serves as a key regulatory node to control pathologic inflammation and suggests that 'metabolic correction' of VLC homeostasis could be an important strategy to normalize dysregulated inflammation caused by the absence of IL-10.
© 2024. The Author(s).
Conflict of interest statement
R.A.F. is an advisor to Glaxo Smith Kline. The other authors declare no competing interests.
Figures





Update of
-
IL-10 constrains sphingolipid metabolism via fatty acid desaturation to limit inflammation.bioRxiv [Preprint]. 2023 May 8:2023.05.07.539780. doi: 10.1101/2023.05.07.539780. bioRxiv. 2023. Update in: Nature. 2024 Mar;627(8004):628-635. doi: 10.1038/s41586-024-07098-5. PMID: 37214856 Free PMC article. Updated. Preprint.

