Identification of hypoxia- and immune-related biomarkers in patients with ischemic stroke
- PMID: 38384585
- PMCID: PMC10878920
- DOI: 10.1016/j.heliyon.2024.e25866
Identification of hypoxia- and immune-related biomarkers in patients with ischemic stroke
Abstract
Background: The immune microenvironment and hypoxia play crucial roles in the pathophysiology of ischemic stroke (IS). Hence, in this study, we aimed to identify hypoxia- and immune-related biomarkers in IS.
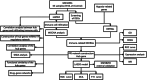
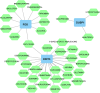
Methods: The IS microarray dataset GSE16561 was examined to determine differentially expressed genes (DEGs) utilizing bioinformatics-based analysis. The intersection of hypoxia-related genes and DEGs was conducted to identify differentially expressed hypoxia-related genes (DEHRGs). Then, using weighted correlation network analysis (WGCNA), all of the genes in GSE16561 dataset were examined to create a co-expression network, and module-clinical trait correlations were examined for the purpose of examining the genes linked to immune cells. The immune-related DEHRGs were submitted to gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses. A protein-protein interaction (PPI) network was constructed by Cytoscape plugin MCODE, in order to extract hub genes. The miRNet was used to predict hub gene-related transcription factors (TFs) and miRNAs. Finally, a diagnostic model was developed by least absolute shrinkage and selection operator (LASSO) logistic regression.
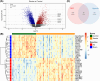
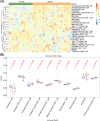
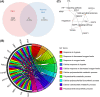
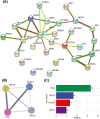
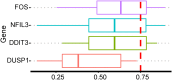
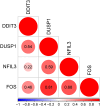
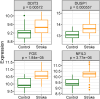
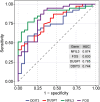
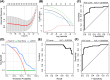
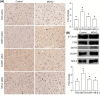
Results: Between the control and IS samples, 4171 DEGs were found. Thereafter, the intersection of hypoxia-related genes and DEGs was conducted to obtain 45 DEHRGs. Ten significantly differentially infiltrated immune cells were found-namely, CD56dim natural killer cells, activated CD8 T cells, activated dendritic cells, activated B cells, central memory CD8 T cells, effector memory CD8 T cells, natural killer cells, gamma delta T cells, plasmacytoid dendritic cells, and neutrophils-between IS and control samples. Subsequently, we identified 27 immune-related DEHRGs through the intersection of DEHRGs and genes in important modules of WGCNA. The immune-related DEHRGs were primarily enriched in response to hypoxia, cellular polysaccharide metabolic process, response to decreased oxygen levels, polysaccharide metabolic process, lipid and atherosclerosis, and HIF-1 signaling pathway H. Using MCODE, FOS, DDIT3, DUSP1, and NFIL3 were found to be hub genes. In the validation cohort and training set, the AUC values of the diagnostic model were 0.9188034 and 0.9395085, respectively.
Conclusion: In brief, we identified and validated four hub genes-FOS, DDIT3, DUSP1, and NFIL3-which might be involved in the pathological development of IS, potentially providing novel perspectives for the diagnosis and treatment of IS.
Keywords: Hub gene; Hypoxia-related genes; Immune cell infiltration; Immune microenvironment; Ischemic stroke.
© 2024 The Authors.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures




Similar articles
-
Identification and validation of oxidative stress and immune-related hub genes in Alzheimer's disease through bioinformatics analysis.Sci Rep. 2023 Jan 12;13(1):657. doi: 10.1038/s41598-023-27977-7. Sci Rep. 2023. PMID: 36635346 Free PMC article.
-
Dysregulation and imbalance of innate and adaptive immunity are involved in the cardiomyopathy progression.Front Cardiovasc Med. 2022 Sep 6;9:973279. doi: 10.3389/fcvm.2022.973279. eCollection 2022. Front Cardiovasc Med. 2022. PMID: 36148059 Free PMC article.
-
Identification of novel biomarkers and immune infiltration characteristics of ischemic stroke based on comprehensive bioinformatic analysis and machine learning.Biochem Biophys Rep. 2023 Dec 7;37:101595. doi: 10.1016/j.bbrep.2023.101595. eCollection 2024 Mar. Biochem Biophys Rep. 2023. PMID: 38371524 Free PMC article.
-
Six potential biomarkers in septic shock: a deep bioinformatics and prospective observational study.Front Immunol. 2023 Jun 8;14:1184700. doi: 10.3389/fimmu.2023.1184700. eCollection 2023. Front Immunol. 2023. PMID: 37359526 Free PMC article.
-
Identification of Effective Diagnostic Biomarkers and Immune Cell Infiltration in Atopic Dermatitis by Comprehensive Bioinformatics Analysis.Front Mol Biosci. 2022 Jul 14;9:917077. doi: 10.3389/fmolb.2022.917077. eCollection 2022. Front Mol Biosci. 2022. PMID: 35911963 Free PMC article.
Cited by
-
Identification of novel inflammatory response-related biomarkers in patients with ischemic stroke based on WGCNA and machine learning.Eur J Med Res. 2025 Mar 22;30(1):195. doi: 10.1186/s40001-025-02454-1. Eur J Med Res. 2025. PMID: 40119397 Free PMC article.
References
-
- Tadi P., Lui F. Acute stroke, in StatPearls. StatPearls Publishing Copyright © 2023, StatPearls Publishing LLC.: Treasure Island (FL) ineligible companies. Disclosure: Forshing Lui declares no relevant financial relationships with ineligible companies. 2023
-
- Katan M., Luft A. Global burden of stroke. Semin Neurol. 2018;38(2):208–211. - PubMed
-
- Amarenco P., et al. Classification of stroke subtypes. Cerebrovasc Dis. 2009;27(5):493–501. - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

