Reassortments in single-stranded DNA multipartite viruses: Confronting expectations based on molecular constraints with field observations
- PMID: 38384786
- PMCID: PMC10880892
- DOI: 10.1093/ve/veae010
Reassortments in single-stranded DNA multipartite viruses: Confronting expectations based on molecular constraints with field observations
Abstract
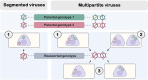
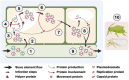
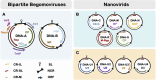
Single-stranded DNA multipartite viruses, which mostly consist of members of the genus Begomovirus, family Geminiviridae, and all members of the family Nanoviridae, partly resolve the cost of genomic integrity maintenance through two remarkable capacities. They are able to systemically infect a host even when their genomic segments are not together in the same host cell, and these segments can be separately transmitted by insect vectors from host to host. These capacities potentially allow such viruses to reassort at a much larger spatial scale, since reassortants could arise from parental genotypes that do not co-infect the same cell or even the same host. To assess the limitations affecting reassortment and their implications in genome integrity maintenance, the objective of this review is to identify putative molecular constraints influencing reassorted segments throughout the infection cycle and to confront expectations based on these constraints with empirical observations. Trans-replication of the reassorted segments emerges as the major constraint, while encapsidation, viral movement, and transmission compatibilities appear more permissive. Confronting the available molecular data and the resulting predictions on reassortments to field population surveys reveals notable discrepancies, particularly a surprising rarity of interspecific natural reassortments within the Nanoviridae family. These apparent discrepancies unveil important knowledge gaps in the biology of ssDNA multipartite viruses and call for further investigation on the role of reassortment in their biology.
Keywords: Begomovirus; Geminiviridae; Nanoviridae; multipartite viruses; reassortment; ssDNA viruses.
© The Author(s) 2024. Published by Oxford University Press.
Conflict of interest statement
None declared.
Figures
Similar articles
-
Insect Transmission of Plant Single-Stranded DNA Viruses.Annu Rev Entomol. 2021 Jan 7;66:389-405. doi: 10.1146/annurev-ento-060920-094531. Epub 2020 Sep 15. Annu Rev Entomol. 2021. PMID: 32931313 Review.
-
Nonconcomitant host-to-host transmission of multipartite virus genome segments may lead to complete genome reconstitution.Proc Natl Acad Sci U S A. 2022 Aug 9;119(32):e2201453119. doi: 10.1073/pnas.2201453119. Epub 2022 Aug 1. Proc Natl Acad Sci U S A. 2022. PMID: 35914138 Free PMC article.
-
Small Bottleneck Size in a Highly Multipartite Virus during a Complete Infection Cycle.J Virol. 2018 Jun 29;92(14):e00139-18. doi: 10.1128/JVI.00139-18. Print 2018 Jul 15. J Virol. 2018. PMID: 29720515 Free PMC article.
-
Replication mechanisms of circular ssDNA plant viruses and their potential implication in viral gene expression regulation.mBio. 2023 Oct 31;14(5):e0169223. doi: 10.1128/mbio.01692-23. Epub 2023 Sep 11. mBio. 2023. PMID: 37695133 Free PMC article. Review.
-
Asymmetric patterns of reassortment and concerted evolution in Cardamom bushy dwarf virus.Infect Genet Evol. 2014 Jun;24:15-24. doi: 10.1016/j.meegid.2014.02.012. Epub 2014 Mar 6. Infect Genet Evol. 2014. PMID: 24613431
Cited by
-
Structural Capsidomics of Single-Stranded DNA Viruses.Viruses. 2025 Feb 27;17(3):333. doi: 10.3390/v17030333. Viruses. 2025. PMID: 40143263 Free PMC article. Review.
References
-
- Alabi O. J. et al. (2021) ‘Olea Europaea Geminivirus Is Present in a Germplasm Repository and in California and Texas Olive (Olea Europaea L.) Groves’, Archives of Virology, 166: 3399–404. - PubMed
-
- Alberter B., Ali Rezaian M., and Jeske H. (2005) ‘Replicative Intermediates of Tomato Leaf Curl Virus and Its Satellite DNAs’, Virology, 331: 441–8. - PubMed
-
- Amin I. et al. (2011) ‘Identification of a Major Pathogenicity Determinant and Suppressors of RNA Silencing Encoded by a South Pacific Isolate of Banana Bunchy Top Virus Originating from Pakistan’, Virus Genes, 42: 272–81. - PubMed
-
- Argüello-Astorga G. R. et al. (1994) ‘Geminivirus Replication Origins Have a Group-specific Organization of Iterative Elements: A Model for Replication’, Virology, 203: 90–100. - PubMed
-
- Argüello-Astorga G. R., and Ruiz-Medrano R. (2001) ‘An Iteron-related Domain Is Associated to Motif 1 in the Replication Proteins of Geminiviruses: Identification of Potential Interacting Amino Acid-Base Pairs by a Comparative Approach’, Archives of Virology, 146: 1465–85. - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Miscellaneous

