TRIM27 elicits protective immunity against tuberculosis by activating TFEB-mediated autophagy flux
- PMID: 38390831
- PMCID: PMC11210901
- DOI: 10.1080/15548627.2024.2321831
TRIM27 elicits protective immunity against tuberculosis by activating TFEB-mediated autophagy flux
Abstract
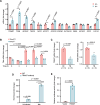
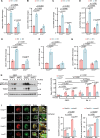
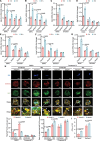
Infectious diseases, such as Mycobacterium tuberculosis (Mtb)-caused tuberculosis (TB), remain a global threat exacerbated by increasing drug resistance. Host-directed therapy (HDT) is a promising strategy for infection treatment through targeting host immunity. However, the limited understanding of the function and regulatory mechanism of host factors involved in immune defense against infections has impeded HDT development. Here, we identify the ubiquitin ligase (E3) TRIM27 (tripartite motif-containing 27) as a host protective factor against Mtb by enhancing host macroautophagy/autophagy flux in an E3 ligase activity-independent manner. Mechanistically, upon Mtb infection, nuclear-localized TRIM27 increases and functions as a transcription activator of TFEB (transcription factor EB). Specifically, TRIM27 binds to the TFEB promoter and the TFEB transcription factor CREB1 (cAMP responsive element binding protein 1), thus enhancing CREB1-TFEB promoter binding affinity and promoting CREB1 transcription activity toward TFEB, eventually inducing autophagy-related gene expression as well as autophagy flux activation to clear the pathogen. Furthermore, TFEB activator 1 can rescue TRIM27 deficiency-caused decreased autophagy-related gene transcription and attenuated autophagy flux, and accordingly suppressed the intracellular survival of Mtb in cell and mouse models. Taken together, our data reveal that TRIM27 is a host defense factor against Mtb, and the TRIM27-CREB1-TFEB axis is a potential HDT-based TB target that can enhance host autophagy flux.Abbreviations: ATG5: autophagy related 5; BMDMs: bone marrow-derived macrophages; CFU: colony-forming unit; ChIP-seq: chromatin immunoprecipitation followed by sequencing; CREB1: cAMP responsive element binding protein 1; CTSB: cathepsin B; E3: ubiquitin ligase; EMSA: electrophoretic mobility shift assay; HC: healthy control; HDT: host-directed therapy; LAMP: lysosomal associated membrane protein; MAP1LC3/LC3: microtubule associated protein 1 light chain 3; MCOLN1: mucolipin TPR cation channel 1; Mtb: Mycobacterium tuberculosis; NLS: nuclear localization signal; PBMCs: peripheral blood mononuclear cells; PRKA/PKA: protein kinase cAMP-activated; qRT-PCR: quantitative real-time PCR; RFP: RET finger protein; TB: tuberculosis; TBK1: TANK binding kinase 1; TFEB: transcription factor EB; TRIM: tripartite motif; TSS: transcription start site; ULK1: unc-51 like autophagy activating kinase 1.
Keywords: Autophagy flux; Mycobacterium tuberculosis; cAMP responsive element binding protein 1; transcription factor EB; tripartite motif-containing 27; tuberculosis.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures






References
-
- World Health Organization . Global Tuberculosis Report 2023. Geneva: World Health Organization; 2023. https://www.who.int/teams/global-tuberculosis-programme/tb-reports/globa....
-
- World Health Organization . Global research agenda for antimicrobial resistance in human health. Geneva: World Health Organization; 2023. https://www.who.int/publications/m/item/global-research-agenda-for-antim....
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous
