A Concise Review of Biomolecule Visualization
- PMID: 38392202
- PMCID: PMC10887528
- DOI: 10.3390/cimb46020084
A Concise Review of Biomolecule Visualization
Abstract
The structural characteristics of biomolecules are a major focus in the field of structural biology. Molecular visualization plays a crucial role in displaying structural information in an intuitive manner, aiding in the understanding of molecular properties. This paper provides a comprehensive overview of core concepts, key techniques, and tools in molecular visualization. Additionally, it presents the latest research findings to uncover emerging trends and highlights the challenges and potential directions for the development of the field.
Keywords: augmented reality; computer graphics; level of detail; molecular visualization; representation models.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures




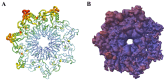
References
-
- Perkins J.A. A history of molecular representation. Part one: 1800 to the 1960s. J. Biocommun. 2005;31:1
-
- Max N.L. ATOMLLL: ATOMS with shading and highlights. ACM SIGGRAPH Comput. Graph. 1979;13:165–173. doi: 10.1145/965103.807439. - DOI
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources

