Augmented interpretation of HER2, ER, and PR in breast cancer by artificial intelligence analyzer: enhancing interobserver agreement through a reader study of 201 cases
- PMID: 38395930
- PMCID: PMC10885430
- DOI: 10.1186/s13058-024-01784-y
Augmented interpretation of HER2, ER, and PR in breast cancer by artificial intelligence analyzer: enhancing interobserver agreement through a reader study of 201 cases
Abstract
Background: Accurate classification of breast cancer molecular subtypes is crucial in determining treatment strategies and predicting clinical outcomes. This classification largely depends on the assessment of human epidermal growth factor receptor 2 (HER2), estrogen receptor (ER), and progesterone receptor (PR) status. However, variability in interpretation among pathologists pose challenges to the accuracy of this classification. This study evaluates the role of artificial intelligence (AI) in enhancing the consistency of these evaluations.
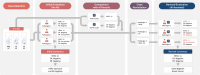
Methods: AI-powered HER2 and ER/PR analyzers, consisting of cell and tissue models, were developed using 1,259 HER2, 744 ER, and 466 PR-stained immunohistochemistry (IHC) whole-slide images of breast cancer. External validation cohort comprising HER2, ER, and PR IHCs of 201 breast cancer cases were analyzed with these AI-powered analyzers. Three board-certified pathologists independently assessed these cases without AI annotation. Then, cases with differing interpretations between pathologists and the AI analyzer were revisited with AI assistance, focusing on evaluating the influence of AI assistance on the concordance among pathologists during the revised evaluation compared to the initial assessment.
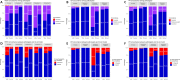
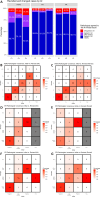
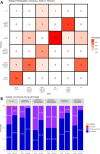
Results: Reevaluation was required in 61 (30.3%), 42 (20.9%), and 80 (39.8%) of HER2, in 15 (7.5%), 17 (8.5%), and 11 (5.5%) of ER, and in 26 (12.9%), 24 (11.9%), and 28 (13.9%) of PR evaluations by the pathologists, respectively. Compared to initial interpretations, the assistance of AI led to a notable increase in the agreement among three pathologists on the status of HER2 (from 49.3 to 74.1%, p < 0.001), ER (from 93.0 to 96.5%, p = 0.096), and PR (from 84.6 to 91.5%, p = 0.006). This improvement was especially evident in cases of HER2 2+ and 1+, where the concordance significantly increased from 46.2 to 68.4% and from 26.5 to 70.7%, respectively. Consequently, a refinement in the classification of breast cancer molecular subtypes (from 58.2 to 78.6%, p < 0.001) was achieved with AI assistance.
Conclusions: This study underscores the significant role of AI analyzers in improving pathologists' concordance in the classification of breast cancer molecular subtypes.
Keywords: Artificial intelligence (AI); Breast cancer; Concordance; Digital pathology; Estrogen receptor (ER); Human epidermal growth factor receptor 2 (HER2); Progesterone receptor (PR); Whole-slide image (WSI).
© 2024. The Author(s).
Conflict of interest statement
Soo Ick Cho, Sangwon Shin, Taebum Lee, Wonkyung Jung, Hajin Lee, Jiyoung Park, Sanghoon Song, Gahee Park, Heon Song, Seonwook Park, Jinhee Lee, Mingu Kang, Jongchan Park, Sergio Pereira, Donggeun Yoo, Keunhyung Chung, and Siraj M. Ali are employees of Lunit and/or have stock/stock options in Lunit.
Figures

Similar articles
-
Systematic Analysis of Factors Affecting Human Epidermal Growth Factor Receptor 2 Interpretation Consistency: Staining Protocols, Artificial Intelligence-Based Image Standardization, and Classification Criteria.Lab Invest. 2025 Jun;105(6):104134. doi: 10.1016/j.labinv.2025.104134. Epub 2025 Mar 23. Lab Invest. 2025. PMID: 40132671
-
Comparison of immunohistochemistry and RT-qPCR for assessing ER, PR, HER2, and Ki67 and evaluating subtypes in patients with breast cancer.Breast Cancer Res Treat. 2022 Aug;194(3):517-529. doi: 10.1007/s10549-022-06649-6. Epub 2022 Jul 5. Breast Cancer Res Treat. 2022. PMID: 35789315
-
Estrogen receptor, progesterone receptor, and human epidermal growth factor receptor 2 expression in breast cancer FNA cell blocks and paired histologic specimens: A large retrospective study.Cancer Cytopathol. 2016 Nov;124(11):828-835. doi: 10.1002/cncy.21745. Epub 2016 Jun 17. Cancer Cytopathol. 2016. PMID: 27315045
-
Insights into AI advances in immunohistochemistry for effective breast cancer treatment: a literature review of ER, PR, and HER2 scoring.Curr Med Res Opin. 2025 Jan;41(1):115-134. doi: 10.1080/03007995.2024.2445142. Epub 2025 Jan 6. Curr Med Res Opin. 2025. PMID: 39705612 Review.
-
Diagnostic performance of ultrasound-based artificial intelligence for predicting key molecular markers in breast cancer: A systematic review and meta-analysis.PLoS One. 2024 May 31;19(5):e0303669. doi: 10.1371/journal.pone.0303669. eCollection 2024. PLoS One. 2024. PMID: 38820391 Free PMC article.
Cited by
-
Corr-A-Net: Interpretable Attention-Based Correlated Feature Learning framework for predicting of HER2 Score in Breast Cancer from H&E Images.medRxiv [Preprint]. 2025 Apr 25:2025.04.22.25326227. doi: 10.1101/2025.04.22.25326227. medRxiv. 2025. PMID: 40313277 Free PMC article. Preprint.
-
Fully automatic HER2 tissue segmentation for interpretable HER2 scoring.J Pathol Inform. 2025 Mar 18;17:100435. doi: 10.1016/j.jpi.2025.100435. eCollection 2025 Apr. J Pathol Inform. 2025. PMID: 40236564 Free PMC article.
-
Interobserver consistency and diagnostic challenges in HER2-ultralow breast cancer: a multicenter study.ESMO Open. 2025 Feb;10(2):104127. doi: 10.1016/j.esmoop.2024.104127. Epub 2025 Jan 31. ESMO Open. 2025. PMID: 39891991 Free PMC article.
-
Semi-automated analysis of HER2 immunohistochemistry in invasive breast carcinoma using whole slide images: utility for interpretation in clinical practice.Pathol Oncol Res. 2024 Aug 29;30:1611826. doi: 10.3389/pore.2024.1611826. eCollection 2024. Pathol Oncol Res. 2024. PMID: 39267995 Free PMC article.
-
A universal immunohistochemistry analyzer for generalizing AI-driven assessment of immunohistochemistry across immunostains and cancer types.NPJ Precis Oncol. 2024 Dec 3;8(1):277. doi: 10.1038/s41698-024-00770-z. NPJ Precis Oncol. 2024. PMID: 39627299 Free PMC article.
References
-
- Goldhirsch A, Winer EP, Coates AS, Gelber RD, Piccart-Gebhart M, Thürlimann B, et al. Personalizing the treatment of women with early breast cancer: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann Oncol. 2013;24:2206–2223. doi: 10.1093/annonc/mdt303. - DOI - PMC - PubMed
-
- Early Breast Cancer Trialists’ Collaborative Group (EBCTCG), Davies C, Godwin J, Gray R, Clarke M, Cutter D, et al. Relevance of breast cancer hormone receptors and other factors to the efficacy of adjuvant tamoxifen: patient-level meta-analysis of randomised trials. Lancet. 2011;378:771–84. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

