Shared and Compartment-Specific Processes in Nucleus Pulposus and Annulus Fibrosus During Intervertebral Disc Degeneration
- PMID: 38403470
- PMCID: PMC11077672
- DOI: 10.1002/advs.202309032
Shared and Compartment-Specific Processes in Nucleus Pulposus and Annulus Fibrosus During Intervertebral Disc Degeneration
Abstract
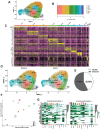
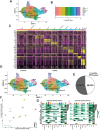
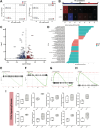
Elucidating how cell populations promote onset and progression of intervertebral disc degeneration (IDD) has the potential to enable more precise therapeutic targeting of cells and mechanisms. Single-cell RNA-sequencing (scRNA-seq) is performed on surgically separated annulus fibrosus (AF) (19,978; 26,983 cells) and nucleus pulposus (NP) (20,884; 24,489 cells) from healthy and diseased human intervertebral discs (IVD). In both tissue types, depletion of cell subsets involved in maintenance of healthy IVD is observed, specifically the immature cell subsets - fibroblast progenitors and stem cells - indicative of an impairment of normal tissue self-renewal. Tissue-specific changes are also identified. In NP, several fibrotic populations are increased in degenerated IVD, indicating tissue-remodeling. In degenerated AF, a novel disease-associated subset is identified, which expresses disease-promoting genes. It is associated with pathogenic biological processes and the main gene regulatory networks include thrombospondin signaling and FOXO1 transcription factor. In NP and AF cells thrombospondin protein promoted expression of genes associated with TGFβ/fibrosis signaling, angiogenesis, and nervous system development. The data reveal new insights of both shared and tissue-specific changes in specific cell populations in AF and NP during IVD degeneration. These identified mechanisms and molecules are novel and more precise targets for IDD prevention and treatment.
Keywords: FOXO transcription factors; intervertebral disc (IVD); intervertebral disc degeneration (IDD); single‐cell RNA‐sequencing (scRNA‐seq); thrombospondin (THBS) signaling.
© 2024 The Authors. Advanced Science published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures







References
-
- Vos T., Flaxman A. D., Naghavi M., Lozano R., Michaud C., Ezzati M., Shibuya K., Salomon J. A., Abdalla S., Aboyans V., Lancet 2012, 380, 2163.
-
- a) Katz J. N., JBJS 2006, 88, 21;
- b) Martin B. I., Deyo R. A., Mirza S. K., Turner J. A., Comstock B. A., Hollingworth W., Sullivan S. D., JAMA, J. Am. Med. Assoc. 2008, 299, 656. - PubMed
-
- Hirsch C., Ingelmark B.‐E., Miller M., Acta Orthop. Scand. 1963, 33, 1. - PubMed
-
- Manchikanti L., Singh V., Falco F. J., Benyamin R. M., Hirsch J. A., Neuromodul.: Technol. Neur. Interface 2014, 17, 3.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous
