This is a preprint.
Integrated transcriptomic analysis of human induced pluripotent stem cell-derived osteogenic differentiation reveals a regulatory role of KLF16
- PMID: 38405902
- PMCID: PMC10888757
- DOI: 10.1101/2024.02.11.579844
Integrated transcriptomic analysis of human induced pluripotent stem cell-derived osteogenic differentiation reveals a regulatory role of KLF16
Abstract
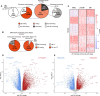
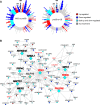
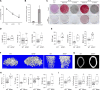
Osteogenic differentiation is essential for bone development, metabolism, and repair; however, the underlying regulatory relationships among genes remain poorly understood. To elucidate the transcriptomic changes and identify novel regulatory genes involved in osteogenic differentiation, we differentiated mesenchymal stem cells (MSCs) derived from 20 human iPSC lines into preosteoblasts (preOBs) and osteoblasts (OBs). We then performed transcriptome profiling of MSCs, preOBs and OBs. The iPSC-derived MSCs and OBs showed similar transcriptome profiles to those of primary human MSCs and OBs, respectively. Differential gene expression analysis revealed global changes in the transcriptomes from MSCs to preOBs, and then to OBs, including the differential expression of 840 genes encoding transcription factors (TFs). TF regulatory network analysis uncovered a network comprising 451 TFs, organized into five interactive modules. Multiscale embedded gene co-expression network analysis (MEGENA) identified gene co-expression modules and key network regulators (KNRs). From these analyses, KLF16 emerged as an important TF in osteogenic differentiation. We demonstrate that overexpression of Klf16 in vitro inhibited osteogenic differentiation and mineralization, while Klf16 +/- mice exhibited increased bone mineral density, trabecular number, and cortical bone area. Our study underscores the complexity of osteogenic differentiation and identifies novel regulatory genes such as KLF16, which plays an inhibitory role in osteogenic differentiation both in vitro and in vivo.
Keywords: RNA sequencing; bone; differential gene expression; induced pluripotent stem cell; mesenchymal stem cell; network analysis; osteoblast; single-cell RNA-seq; systems biology; transcription factor.
Figures


References
-
- Al-Rekabi Z., Wheeler M. M., Leonard A., Fura A. M., Juhlin I., Frazar C., Smith J. D., Park S. S., Gustafson J. A., Clarke C. M., et al. (2016). Activation of the IGF1 pathway mediates changes in cellular contractility and motility in single-suture craniosynostosis. Journal of Cell Science 129, 483–491. - PMC - PubMed
-
- Bastian M., Heymann S. and Jacomy M. (2009). Gephi: an open source software for exploring and manipulating networks. Proceedings of the International AAAI Conference on Web and Social Media 3, 361–362.
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
