Mechanistic insights into the alternative ribosome recycling by HflXr
- PMID: 38407413
- PMCID: PMC11040002
- DOI: 10.1093/nar/gkae128
Mechanistic insights into the alternative ribosome recycling by HflXr
Erratum in
-
Correction to 'Mechanistic insights into the alternative ribosome recycling by HflXr'.Nucleic Acids Res. 2024 Apr 24;52(7):4109. doi: 10.1093/nar/gkae217. Nucleic Acids Res. 2024. PMID: 38499499 Free PMC article. No abstract available.
Abstract
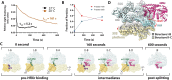
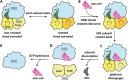
During stress conditions such as heat shock and antibiotic exposure, ribosomes stall on messenger RNAs, leading to inhibition of protein synthesis. To remobilize ribosomes, bacteria use rescue factors such as HflXr, a homolog of the conserved housekeeping GTPase HflX that catalyzes the dissociation of translationally inactive ribosomes into individual subunits. Here we use time-resolved cryo-electron microscopy to elucidate the mechanism of ribosome recycling by Listeria monocytogenes HflXr. Within the 70S ribosome, HflXr displaces helix H69 of the 50S subunit and induces long-range movements of the platform domain of the 30S subunit, disrupting inter-subunit bridges B2b, B2c, B4, B7a and B7b. Our findings unveil a unique ribosome recycling strategy by HflXr which is distinct from that mediated by RRF and EF-G. The resemblance between HflXr and housekeeping HflX suggests that the alternative ribosome recycling mechanism reported here is universal in the prokaryotic kingdom.
© The Author(s) 2024. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures





References
-
- Park J.H., Jensen B.C., Kifer C.T., Parsons M. A novel nucleolar G-protein conserved in eukaryotes. J. Cell Sci. 2001; 114:173–185. - PubMed
-
- Brown E.D. Conserved P-loop GTPases of unknown function in bacteria: an emerging and vital ensemble in bacterial physiology. Biochem. Cell Biol. 2005; 83:738–746. - PubMed
-
- Sengupta S., Mondal A., Dutta D., Parrack P. HflX protein protects Escherichia coli from manganese stress. J. Biosci. 2018; 43:1001–1013. - PubMed
-
- Zhang Y., Mandava C.S., Cao W., Li X., Zhang D., Li N., Zhang Y., Zhang X., Qin Y., Mi K. et al. . HflX is a ribosome-splitting factor rescuing stalled ribosomes under stress conditions. Nat. Struct. Mol. Biol. 2015; 22:906–913. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

