TRIM72 restricts lyssavirus infection by inducing K48-linked ubiquitination and proteasome degradation of the matrix protein
- PMID: 38408103
- PMCID: PMC10919858
- DOI: 10.1371/journal.ppat.1011718
TRIM72 restricts lyssavirus infection by inducing K48-linked ubiquitination and proteasome degradation of the matrix protein
Abstract
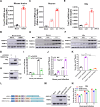
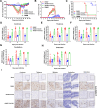
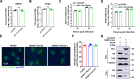
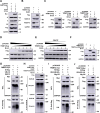
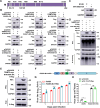
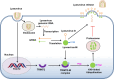
The tripartite motif (TRIM) protein family is the largest subfamily of E3 ubiquitin ligases, playing a crucial role in the antiviral process. In this study, we found that TRIM72, a member of the TRIM protein family, was increased in neuronal cells and mouse brains following rabies lyssavirus (RABV) infection. Over-expression of TRIM72 significantly reduced the viral titer of RABV in neuronal cells and mitigated the pathogenicity of RABV in mice. Furthermore, we found that TRIM72 over-expression effectively prevents the assembly and/or release of RABV. In terms of the mechanism, TRIM72 promotes the K48-linked ubiquitination of RABV Matrix protein (M), leading to the degradation of M through the proteasome pathway. TRIM72 directly interacts with M and the interaction sites were identified and confirmed through TRIM72-M interaction model construction and mutation analysis. Further investigation revealed that the degradation of M induced by TRIM72 was attributed to TRIM72's promotion of ubiquitination at site K195 in M. Importantly, the K195 site was found to be partially conserved among lyssavirus's M proteins, and TRIM72 over-expression induced the degradation of these lyssavirus M proteins. In summary, our study has uncovered a TRIM family protein, TRIM72, that can restrict lyssavirus replication by degrading M, and we have identified a novel ubiquitination site (K195) in lyssavirus M.
Copyright: © 2024 Sui et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
TRIM21 Promotes Rabies Virus Production by Degrading IRF7 through Ubiquitination.Int J Mol Sci. 2023 Jun 30;24(13):10892. doi: 10.3390/ijms241310892. Int J Mol Sci. 2023. PMID: 37446070 Free PMC article.
-
Structure and activation of the RING E3 ubiquitin ligase TRIM72 on the membrane.Nat Struct Mol Biol. 2023 Nov;30(11):1695-1706. doi: 10.1038/s41594-023-01111-7. Epub 2023 Sep 28. Nat Struct Mol Biol. 2023. PMID: 37770719 Free PMC article.
-
RTP4 restricts lyssavirus rabies infection by binding to viral genomic RNA.Vet Microbiol. 2024 Aug;295:110159. doi: 10.1016/j.vetmic.2024.110159. Epub 2024 Jun 19. Vet Microbiol. 2024. PMID: 38941768
-
Emerging Role of TRIM Family Proteins in Cardiovascular Disease.Cardiology. 2020;145(6):390-400. doi: 10.1159/000506150. Epub 2020 Apr 17. Cardiology. 2020. PMID: 32305978 Review.
-
The role of TRIM family proteins in autophagy, pyroptosis, and diabetes mellitus.Cell Biol Int. 2021 May;45(5):913-926. doi: 10.1002/cbin.11550. Epub 2021 Jan 28. Cell Biol Int. 2021. PMID: 33438267 Review.
Cited by
-
Trim72 is a major host factor protecting against lethal Candida albicans infection.PLoS Pathog. 2024 Nov 25;20(11):e1012747. doi: 10.1371/journal.ppat.1012747. eCollection 2024 Nov. PLoS Pathog. 2024. PMID: 39585917 Free PMC article.
-
TRIM proteins: A 'swiss army knife' of antiviral immunity.PLoS Pathog. 2025 May 12;21(5):e1013147. doi: 10.1371/journal.ppat.1013147. eCollection 2025 May. PLoS Pathog. 2025. PMID: 40354393 Free PMC article. Review.
References
-
- Rupprecht CE, Mshelbwala PP, Reeves RG, Kuzmin IV. Rabies in a postpandemic world: resilient reservoirs, redoubtable riposte, recurrent roadblocks, and resolute recidivism. Animal diseases. 2023;3(1):15. Epub 2023/05/30. doi: 10.1186/s44149-023-00078-8 ; PubMed Central PMCID: PMC10195671. - DOI - PMC - PubMed
-
- Hampson K, Coudeville L, Lembo T, Sambo M, Kieffer A, Attlan M, et al.. Estimating the global burden of endemic canine rabies. PLoS neglected tropical diseases. 2015;9(4):e0003709. Epub 2015/04/17. doi: 10.1371/journal.pntd.0003709 ; PubMed Central PMCID: PMC4400070 does not alter our adherence to all PLOS NTDs policies on sharing data and materials. The findings and conclusions in this report are those of the authors and do not necessarily represent the views of the Centers for Disease Control and Prevention. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

