Expression of endogenous epitope-tagged GPR4 in the mouse brain
- PMID: 38408869
- PMCID: PMC10928412
- DOI: 10.1523/ENEURO.0002-24.2024
Expression of endogenous epitope-tagged GPR4 in the mouse brain
Abstract
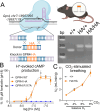
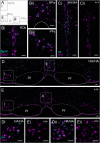
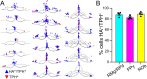
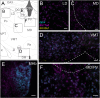
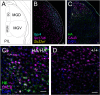
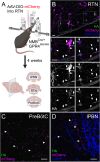
GPR4 is a proton-sensing G protein-coupled receptor implicated in many peripheral and central physiological processes. GPR4 expression has previously been assessed only via detection of the cognate transcript or indirectly, by use of fluorescent reporters. In this work, CRISPR/Cas9 knock-in technology was used to encode a hemagglutinin (HA) epitope tag within the endogenous locus of Gpr4 and visualize GPR4-HA in the mouse central nervous system using a specific, well characterized HA antibody; GPR4 expression was further verified by complementary Gpr4 mRNA detection. HA immunoreactivity was found in a limited set of brain regions, including in the retrotrapezoid nucleus (RTN), serotonergic raphe nuclei, medial habenula, lateral septum, and several thalamic nuclei. GPR4 expression was not restricted to cells of a specific neurochemical identity as it was observed in excitatory, inhibitory, and aminergic neuronal cell groups. HA immunoreactivity was not detected in brain vascular endothelium, despite clear expression of Gpr4 mRNA in endothelial cells. In the RTN, GPR4 expression was detected at the soma and in proximal dendrites along blood vessels and the ventral surface of the brainstem; HA immunoreactivity was not detected in RTN projections to two known target regions. This localization of GPR4 protein in mouse brain neurons corroborates putative sites of expression where its function has been previously implicated (e.g., CO2-regulated breathing by RTN), and provides a guide for where GPR4 could contribute to other CO2/H+ modulated brain functions. Finally, GPR4-HA animals provide a useful reagent for further study of GPR4 in other physiological processes outside of the brain.Significance Statement GPR4 is a proton-sensing G-protein coupled receptor whose expression is necessary for a number of diverse physiological processes including acid-base sensing in the kidney, immune function, and cancer progression. In the brain, GPR4 has been implicated in the hypercapnic ventilatory response mediated by brainstem neurons. While knockout studies in animals have clearly demonstrated its necessity for normal physiology, descriptions of GPR4 expression have been limited due to a lack of specific antibodies for use in mouse models. In this paper, we implemented a CRISPR/Cas9 knock-in approach to incorporate the coding sequence for a small epitope tag into the locus of GPR4. Using these mice, we were able to describe GPR4 protein expression directly for the first time.
Copyright © 2024 Gonye et al.
Conflict of interest statement
The authors declare no competing financial interest.
Figures





References
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
