Gli1 labels progenitors during chondrogenesis in postnatal mice
- PMID: 38409269
- PMCID: PMC11014955
- DOI: 10.1038/s44319-024-00093-x
Gli1 labels progenitors during chondrogenesis in postnatal mice
Abstract
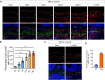
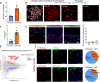
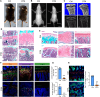
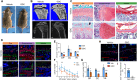
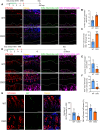
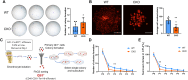
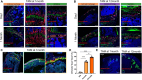
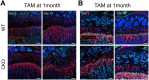
Skeletal growth promoted by endochondral ossification is tightly coordinated by self-renewal and differentiation of chondrogenic progenitors. Emerging evidence has shown that multiple skeletal stem cells (SSCs) participate in cartilage formation. However, as yet, no study has reported the existence of common long-lasting chondrogenic progenitors in various types of cartilage. Here, we identify Gli1+ chondrogenic progenitors (Gli1+ CPs), which are distinct from PTHrP+ or FoxA2+ SSCs, are responsible for the lifelong generation of chondrocytes in the growth plate, vertebrae, ribs, and other cartilage. The absence of Gli1+ CPs leads to cartilage defects and dwarfishness phenotype in mice. Furthermore, we show that the BMP signal plays an important role in self-renewal and maintenance of Gli1+ CPs. Deletion of Bmpr1α triggers Gli1+ CPs quiescence exit and causes the exhaustion of Gli1+ CPs, consequently disrupting columnar cartilage. Collectively, our data demonstrate that Gli1+ CPs are common long-term chondrogenic progenitors in multiple types of cartilage and are essential to maintain cartilage homeostasis.
Keywords: BMP; Chondrogenic Progenitors; Gli1+ Cells; Hedgehog; Stem Cell Maintenance.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



References
-
- Andl T, Ahn K, Kairo A, Chu EY, Wine-Lee L, Reddy ST, Croft NJ, Cebra-Thomas JA, Metzger D, Chambon P, et al. Epithelial Bmpr1a regulates differentiation and proliferation in postnatal hair follicles and is essential for tooth development. Development. 2004;131:2257–2268. doi: 10.1242/dev.01125. - DOI - PubMed
-
- Chagin AS, Vuppalapati KK, Kobayashi T, Guo J, Hirai T, Chen M, Offermanns S, Weinstein LS, Kronenberg HM. G-protein stimulatory subunit alpha and Gq/11alpha G-proteins are both required to maintain quiescent stem-like chondrocytes. Nat Commun. 2014;5:3673. doi: 10.1038/ncomms4673. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
- 81825005/MOST | NSFC | National Science Fund for Distinguished Young Scholars (National Science Foundation for Distinguished Young Scholars)
- NSFC82071091/MOST | National Natural Science Foundation of China (NSFC)
- NSFC82001040/MOST | National Natural Science Foundation of China (NSFC)
- 2020M673265/China Postdoctoral Foundation Project | Postdoctoral Research Foundation of China (China Postdoctoral Research Foundation)
- 2023NSFSC0563/Sichuan Science and Technology Program
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

