Transient disome complex formation in native polysomes during ongoing protein synthesis captured by cryo-EM
- PMID: 38409277
- PMCID: PMC10897467
- DOI: 10.1038/s41467-024-46092-3
Transient disome complex formation in native polysomes during ongoing protein synthesis captured by cryo-EM
Abstract
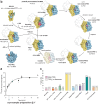
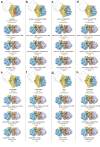
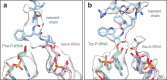
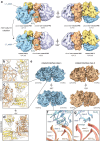
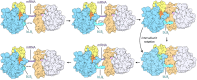
Structural studies of translating ribosomes traditionally rely on in vitro assembly and stalling of ribosomes in defined states. To comprehensively visualize bacterial translation, we reactivated ex vivo-derived E. coli polysomes in the PURE in vitro translation system and analyzed the actively elongating polysomes by cryo-EM. We find that 31% of 70S ribosomes assemble into disome complexes that represent eight distinct functional states including decoding and termination intermediates, and a pre-nucleophilic attack state. The functional diversity of disome complexes together with RNase digest experiments suggests that paused disome complexes transiently form during ongoing elongation. Structural analysis revealed five disome interfaces between leading and queueing ribosomes that undergo rearrangements as the leading ribosome traverses through the elongation cycle. Our findings reveal at the molecular level how bL9's CTD obstructs the factor binding site of queueing ribosomes to thwart harmful collisions and illustrate how translation dynamics reshape inter-ribosomal contacts.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Visualizing translation dynamics at atomic detail inside a bacterial cell.Nature. 2022 Oct;610(7930):205-211. doi: 10.1038/s41586-022-05255-2. Epub 2022 Sep 28. Nature. 2022. PMID: 36171285 Free PMC article.
-
Disome-seq reveals widespread ribosome collisions that promote cotranslational protein folding.Genome Biol. 2021 Jan 5;22(1):16. doi: 10.1186/s13059-020-02256-0. Genome Biol. 2021. PMID: 33402206 Free PMC article.
-
Mechanistic insights into the alternative translation termination by ArfA and RF2.Nature. 2017 Jan 26;541(7638):550-553. doi: 10.1038/nature20822. Epub 2016 Dec 1. Nature. 2017. PMID: 27906160
-
The Structural Dynamics of Translation.Annu Rev Biochem. 2022 Jun 21;91:245-267. doi: 10.1146/annurev-biochem-071921-122857. Epub 2022 Mar 14. Annu Rev Biochem. 2022. PMID: 35287473 Free PMC article. Review.
-
Almost lost in translation. Cryo-EM of a dynamic macromolecular complex: the ribosome.Eur Biophys J. 2011 May;40(5):589-97. doi: 10.1007/s00249-011-0683-6. Epub 2011 Feb 19. Eur Biophys J. 2011. PMID: 21336521 Review.
Cited by
-
Comprehensive translational profiling and STE AI uncover rapid control of protein biosynthesis during cell stress.Nucleic Acids Res. 2024 Jul 22;52(13):7925-7946. doi: 10.1093/nar/gkae365. Nucleic Acids Res. 2024. PMID: 38721779 Free PMC article.
-
The RNA helicase HrpA rescues collided ribosomes in E. coli.Mol Cell. 2025 Mar 6;85(5):999-1007.e7. doi: 10.1016/j.molcel.2025.01.018. Epub 2025 Feb 7. Mol Cell. 2025. PMID: 39922193
-
The RNA helicase HrpA rescues collided ribosomes in E. coli.bioRxiv [Preprint]. 2024 Sep 11:2024.09.11.612461. doi: 10.1101/2024.09.11.612461. bioRxiv. 2024. Update in: Mol Cell. 2025 Mar 06;85(5):999-1007.e7. doi: 10.1016/j.molcel.2025.01.018. PMID: 39314269 Free PMC article. Updated. Preprint.
-
Novel archaeal ribosome dimerization factor facilitating unique 30S-30S dimerization.Nucleic Acids Res. 2025 Jan 11;53(2):gkae1324. doi: 10.1093/nar/gkae1324. Nucleic Acids Res. 2025. PMID: 39797736 Free PMC article.
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

