Validation of a real-time polymerase chain reaction for the detection and quantification of the nucleic acid of Histoplasma from equine clinical samples
- PMID: 38411051
- PMCID: PMC10986522
- DOI: 10.1128/spectrum.03100-23
Validation of a real-time polymerase chain reaction for the detection and quantification of the nucleic acid of Histoplasma from equine clinical samples
Abstract
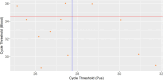
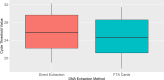
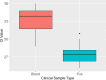
Histoplasma capsulatum var. farciminosum (HCF) is a dimorphic fungus that causes epizootic lymphangitis in equids. Current diagnostic approaches, including culture, microscopy, and clinical presentation, lack speed, sensitivity, and specificity when diagnosing clinical cases. In this study, equine blood and pus samples on Whatman FTA cards from Senegal (n = 3), The Gambia (n = 19), Ethiopia (n = 16), and Mali (n = 13) were tested using a real-time PCR (qPCR) protocol. The assay was optimized and tested for its suitability to detect and quantify HCF in blood and pus loaded onto Whatman FTA cards at sampling. Whatman FTA cards were tested for their suitability for use with qPCR and were found to recover DNA more efficiently than from direct extraction. Using TaqMan fluorescent probes and specific primers, the assay demonstrated 100% analytical specificity when detecting multiple strains of Histoplasma and no false positives with off-target organisms. The assay's diagnostic performance was measured against an existing nested internal transcribed spacer PCR protocol using a receiver operating characteristic curve. The test was found to have a diagnostic specificity and sensitivity of 100% and 71.4%, respectively, when analyzing pus samples using a cycle threshold (Ct) cutoff determined by Youden's index (27.75). Blood sample cutoff Ct value was proposed at 34.55. Further optimization is required to improve the performance of the protocol when applied to blood samples. This study has, for the first time, demonstrated the ability to detect and quantify the DNA of Histoplasma spp. in equine blood and pus samples with a high degree of accuracy, providing a platform to further investigate the pathogenesis and epidemiology of this disease.
Importance: Histoplasmosis is a neglected yet major cause of morbidity and mortality in both equids and people in resource-scarce settings. One of the major hindrances to the control of histoplasmosis is a lack of readily available diagnostic tests. Tests are needed to support clinical decision-making and to be applied in population-based research to further understand this disease in situ. This paper reports, for the first time, the validation and application of a qPCR to detect Histoplasma directly from equine clinical samples, bypassing the need to culture this notoriously difficult organism. We report and comment on the performance of the qPCR in comparison with our previously developed nested PCR.
Keywords: Histoplasma and West Africa; detecting Histoplasma equine samples; equine clinical samples; qPCR Histoplasma.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





References
-
- Endebu B, Roger F. 2003. Comparative studies on the occurrence and distribution of epizootic lymphangitis and ulcerative lymphangitis in Ethiopia. Int J Appl Res Vet Med 1:219–224.
-
- WAHID Interface OIE . 2005. Epizootic lymphangitis disease distribution map. Available from: http://www.oie.int/wahis_2/public/wahid.php/Diseaseinformation/Diseasedi...
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

