The emergence of circadian timekeeping in the intestine
- PMID: 38413599
- PMCID: PMC10899604
- DOI: 10.1038/s41467-024-45942-4
The emergence of circadian timekeeping in the intestine
Abstract
The circadian clock is a molecular timekeeper, present from cyanobacteria to mammals, that coordinates internal physiology with the external environment. The clock has a 24-h period however development proceeds with its own timing, raising the question of how these interact. Using the intestine of Drosophila melanogaster as a model for organ development, we track how and when the circadian clock emerges in specific cell types. We find that the circadian clock begins abruptly in the adult intestine and gradually synchronizes to the environment after intestinal development is complete. This delayed start occurs because individual cells at earlier stages lack the complete circadian clock gene network. As the intestine develops, the circadian clock is first consolidated in intestinal stem cells with changes in Ecdysone and Hnf4 signalling influencing the transcriptional activity of Clk/cyc to drive the expression of tim, Pdp1, and vri. In the mature intestine, stem cell lineage commitment transiently disrupts clock activity in differentiating progeny, mirroring early developmental clock-less transitions. Our data show that clock function and differentiation are incompatible and provide a paradigm for studying circadian clocks in development and stem cell lineages.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




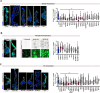


References
MeSH terms
Substances
Grants and funding
- RGPIN-2020-04252/Canadian Network for Research and Innovation in Machining Technology, Natural Sciences and Engineering Research Council of Canada (NSERC Canadian Network for Research and Innovation in Machining Technology)
- RGPIN-2023-04028/Gouvernement du Canada | Natural Sciences and Engineering Research Council of Canada (Conseil de Recherches en Sciences Naturelles et en Génie du Canada)
- MOP77746/Gouvernement du Canada | Canadian Institutes of Health Research (Instituts de Recherche en Santé du Canada)
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

