A 5.2-kb insertion in the coding sequence of PavSCPL, a serine carboxypeptidase-like enhances fruit firmness in Prunus avium
- PMID: 38415985
- PMCID: PMC11123409
- DOI: 10.1111/pbi.14291
A 5.2-kb insertion in the coding sequence of PavSCPL, a serine carboxypeptidase-like enhances fruit firmness in Prunus avium
Abstract
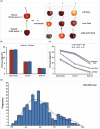
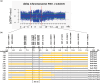
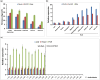
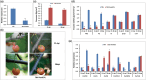
Fruit firmness is an important trait in sweet cherry breeding because it directly positively influences fruit transportability, storage and shelf life. However, the underlying genes responsible and the molecular mechanisms that control fruit firmness remain unknown. In this study, we identified a candidate gene, PavSCPL, encoding a serine carboxypeptidase-like protein with natural allelic variation, that controls fruit firmness in sweet cherry using map-based cloning and functionally characterized PavSCPL during sweet cherry fruit softening. Genetic analysis revealed that fruit firmness in the 'Rainier' × 'Summit' F1 population was controlled by a single dominant gene. Bulked segregant analysis combined with fine mapping narrowed the candidate gene to a 473-kb region (7418778-7 891 914 bp) on chromosome 6 which included 72 genes. The candidate gene PavSCPL, and a null allele harbouring a 5244-bp insertion in the second exon that completely inactivated PavSCPL expression and resulted in the extra-hard-flesh phenotype, were identified by RNA-sequencing analysis and gene cloning. Quantitative RT-PCR analysis revealed that the PavSCPL expression level was increased with fruit softening. Virus-induced gene silencing of PavSCPL enhanced fruit firmness and suppressed the activities of certain pectin-degrading enzymes in the fruit. In addition, we developed functional molecular markers for PavSCPL and the Pavscpl5.2-k allele that co-segregated with the fruit firmness trait. Overall, this research identified a crucial functional gene for fruit firmness. The results provide insights into the genetic control and molecular mechanism of the fruit firmness trait and present useful molecular markers for molecular-assisted breeding for fruit firmness in sweet cherry.
Keywords: BSA‐seq; PavSCPL; fruit firmness; molecular marker; sweet cherry.
© 2024 The Authors. Plant Biotechnology Journal published by Society for Experimental Biology and The Association of Applied Biologists and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare that they have no conflicts of interest.
Figures


References
-
- Ahmad, M.Z. , Li, P. , She, G. , Xia, E. , Benedito, V.A. , Wan, X.C. and Zhao, J. (2020) Genome‐wide analysis of serine carboxypeptidase‐like acyltransferase gene family for evolution and characterization of enzymes involved in the biosynthesis of galloylated catechins in the tea plant (Camellia sinensis). Front. Plant Sci. 11, 848. - PMC - PubMed
-
- Bashline, L. , Lei, L. , Li, S. and Gu, Y. (2014) Cell wall, cytoskeleton, and cell expansion in higher plants. Mol. Plant, 7, 586–600. - PubMed
-
- Brummell, D.A. and Harpster, M.H. (2001) Cell wall metabolism in fruit softening and quality and its manipulation in transgenic plants. Plant Mol. Biol. 47, 311–340. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

