Fibroblast Growth Factor 19 Alters Bile Acids to Induce Dysbiosis in Mice With Alcohol-Induced Liver Disease
- PMID: 38417701
- PMCID: PMC11127034
- DOI: 10.1016/j.jcmgh.2024.02.015
Fibroblast Growth Factor 19 Alters Bile Acids to Induce Dysbiosis in Mice With Alcohol-Induced Liver Disease
Abstract
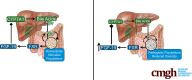
Background & aims: Excessive alcohol consumption can lead to alcohol-associated liver disease, a spectrum of conditions ranging from steatosis to fibrosis and cirrhosis. Bile acids regulate metabolic pathways by binding to cellular and nuclear receptors, and they also interact with the gut microbiome to control microbial overgrowth. Fibroblast growth factor 19 (FGF-19) is an ileum-derived hormone induced and released in response to bile acid activation of the nuclear receptor farnesoid X receptor. FGF-19 signaling is dysregulated with ethanol consumption and is increased in patients with alcoholic hepatitis. Here, we examined the effects of FGF-19 in a mouse model of chronic + binge ethanol feeding.
Methods: After injection of adeno-associated virus-green fluorescent protein or AAV-FGF-19, female C57BL/6J mice were pair-fed a Lieber DeCarli liquid diet (5% v/v) or control diet for 10 days and were given a bolus gavage of 5% ethanol or maltose control to represent a binge drinking episode. Tissues were collected for analysis 9 hours after the binge.
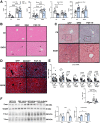
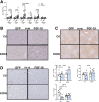
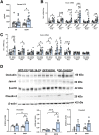
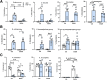
Results: Chronic + binge ethanol feeding induced steatosis regardless of FGF-19 expression. Interestingly, FGF-19 and ethanol resulted in significantly increased liver inflammation, as measured by Il6, Tgfβ, and Tnfα, compared with ethanol alone. Both ethanol and FGF-19 decreased bile acid synthesis, and FGF-19 significantly reduced secondary bile acids, leading to overgrowth of specific pathogenic bacteria including Enterococcus faecalis, Escherichia coli, and Clostridium perfringens.
Conclusions: Dysregulation of FGF-19 and consequent changes in bile acid synthesis and composition during alcohol consumption may be a contributing factor to alcohol-induced liver disease and dysbiosis.
Keywords: Ethanol; Inflammation; Microbiome.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Figures




Similar articles
-
Apolipoprotein H deficiency exacerbates alcohol-induced liver injury via gut Dysbiosis and altered bile acid metabolism.Biochim Biophys Acta Mol Cell Biol Lipids. 2024 Oct;1869(7):159535. doi: 10.1016/j.bbalip.2024.159535. Epub 2024 Jul 20. Biochim Biophys Acta Mol Cell Biol Lipids. 2024. PMID: 39033850
-
Use of a crossed high alcohol preferring (cHAP) mouse model with the NIAAA-model of chronic-binge ethanol intake to study liver injury.Alcohol Alcohol. 2017 Nov 1;52(6):629-637. doi: 10.1093/alcalc/agx063. Alcohol Alcohol. 2017. PMID: 29036399 Free PMC article.
-
The selective PPAR-delta agonist seladelpar reduces ethanol-induced liver disease by restoring gut barrier function and bile acid homeostasis in mice.Transl Res. 2021 Jan;227:1-14. doi: 10.1016/j.trsl.2020.06.006. Epub 2020 Jun 15. Transl Res. 2021. PMID: 32553670 Free PMC article.
-
Rodent Models of Alcoholic Liver Disease: Role of Binge Ethanol Administration.Biomolecules. 2018 Jan 13;8(1):3. doi: 10.3390/biom8010003. Biomolecules. 2018. PMID: 29342874 Free PMC article. Review.
-
Impact of binge drinking on alcoholic liver disease.Arch Pharm Res. 2025 Mar;48(3):212-223. doi: 10.1007/s12272-025-01537-1. Epub 2025 Mar 4. Arch Pharm Res. 2025. PMID: 40035998 Review.
Cited by
-
Interorgan and Transcellular Communication in Metabolic Disease: Insights From Recent CMGH Studies.Cell Mol Gastroenterol Hepatol. 2025;19(8):101530. doi: 10.1016/j.jcmgh.2025.101530. Epub 2025 May 8. Cell Mol Gastroenterol Hepatol. 2025. PMID: 40348017 Free PMC article. Review.
-
Transcellular, Interorgan, and Host-microbial Communication in Metabolic Homeostasis: Hearing for the First Time What We Once Thought to be Unspoken.Cell Mol Gastroenterol Hepatol. 2025;19(9):101545. doi: 10.1016/j.jcmgh.2025.101545. Epub 2025 Jun 9. Cell Mol Gastroenterol Hepatol. 2025. PMID: 40532271 Free PMC article. No abstract available.
-
Fibroblast Growth Factor 19 in Alcohol-Associated Liver Disease: Bile Acids and Dysbiosis and Inflammation, Oh My!Cell Mol Gastroenterol Hepatol. 2024;18(1):159-160. doi: 10.1016/j.jcmgh.2024.03.014. Epub 2024 Apr 15. Cell Mol Gastroenterol Hepatol. 2024. PMID: 38636549 Free PMC article. No abstract available.
-
The Impact of Human Liver Transplantation on the Concentration of Fibroblast Growth Factors: FGF19 and FGF21.Int J Mol Sci. 2025 Feb 3;26(3):1299. doi: 10.3390/ijms26031299. Int J Mol Sci. 2025. PMID: 39941067 Free PMC article.
References
-
- Carvalho A.F., Heilig M., Perez A., et al. Alcohol use disorders. Lancet. 2019;394:781–792. - PubMed
-
- Kezer C.A., Simonetto D.A., Shah V.H. Sex differences in alcohol consumption and alcohol-associated liver disease. Mayo Clin Proc. 2021;96:1006–1016. - PubMed
-
- Mellinger J.L., Shedden K., Winder G.S., et al. The high burden of alcoholic cirrhosis in privately insured persons in the United States. Hepatology. 2018;68:872–882. - PubMed
-
- Wong R.J., Aguilar M., Cheung R., et al. Nonalcoholic steatohepatitis is the second leading etiology of liver disease among adults awaiting liver transplantation in the United States. Gastroenterology. 2015;148:547–555. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

