Comparison of pre-labelled primers and nucleotides as DNA labelling method for lateral flow detection of Legionella pneumophila amplicons
- PMID: 38424185
- PMCID: PMC10904838
- DOI: 10.1038/s41598-024-55703-4
Comparison of pre-labelled primers and nucleotides as DNA labelling method for lateral flow detection of Legionella pneumophila amplicons
Abstract
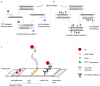
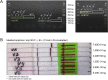
Labelling of nucleic acid amplicons during polymerase chain reaction (PCR) or isothermal techniques is possible by using both labelled primers and labelled nucleotides. While the former is the widely used method, the latter can offer significant advantages in terms of signal enhancement and improving the detection limit of an assay. Advantages and disadvantages of both methods depend on different factors, including amplification method, detection method and amplicon length. In this study, both methods for labelling PCR products for lateral flow assay (LFA) analysis (LFA-PCR) were analysed and compared. It was shown that labelling by means of nucleotides results in an increase in label incorporation rates. Nonetheless, this advantage is negated by the need for post-processing and competitive interactions. In the end, it was possible to achieve a detection limit of 3 cell equivalents for the detection of the Legionella-DNA used here via primer labelling. Labelling via nucleotides required genomic DNA of at least 3000 cell equivalents as starting material as well as an increased personnel and experimental effort.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
[Establishment of loop-mediated isothermal amplification method for detection of Legionella pneumophila].Zhejiang Da Xue Xue Bao Yi Xue Ban. 2010 May;39(3):305-10. doi: 10.3785/j.issn.1008-9292.2010.03.015. Zhejiang Da Xue Xue Bao Yi Xue Ban. 2010. PMID: 20544994 Chinese.
-
Investigation and validation of labelling loop mediated isothermal amplification (LAMP) products with different nucleotide modifications for various downstream analysis.Sci Rep. 2022 May 3;12(1):7137. doi: 10.1038/s41598-022-11320-7. Sci Rep. 2022. PMID: 35504953 Free PMC article.
-
[Establishment and application of loop-mediated isothermal amplification method for rapid detection].Zhonghua Liu Xing Bing Xue Za Zhi. 2009 May;30(5):481-5. Zhonghua Liu Xing Bing Xue Za Zhi. 2009. PMID: 19799145 Chinese.
-
On-filter direct amplification of Legionella pneumophila for rapid assessment of its abundance and viability.Water Res. 2017 Sep 15;121:162-170. doi: 10.1016/j.watres.2017.05.028. Epub 2017 May 13. Water Res. 2017. PMID: 28527390 Free PMC article.
-
Loop-mediated isothermal amplification of DNA (LAMP): a new diagnostic tool lights the world of diagnosis of animal and human pathogens: a review.Pak J Biol Sci. 2014 Jan 15;17(2):151-66. doi: 10.3923/pjbs.2014.151.166. Pak J Biol Sci. 2014. PMID: 24783797 Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

