Expression of PTGS2 along with genes regulating VEGF signalling pathway and association with high-risk factors in locally advanced oral squamous cell carcinoma
- PMID: 38426619
- PMCID: PMC10905678
- DOI: 10.1002/cam4.6986
Expression of PTGS2 along with genes regulating VEGF signalling pathway and association with high-risk factors in locally advanced oral squamous cell carcinoma
Abstract
Background: PTGS2 encodes cyclooxygenase-2 (COX-2), which catalyses the committed step in prostaglandin synthesis. Various in vivo and in vitro data suggest that COX-2 mediates the VEGF signalling pathway. In silico analysis performed in TCGA, PanCancer Atlas for head and neck cancers, demonstrated significant expression and co-expression of PTGS2 and genes that regulate VEGF signalling. This study was designed to elucidate the expression pattern of PTGS2 and genes regulating VEGF signalling in patients with locally advanced oral squamous cell carcinoma (OSCC).
Methodology: Tumour and normal tissue samples were collected from patients with locally advanced OSCC. RNA was isolated from tissue samples, followed by cDNA synthesis. The cDNA was used for gene expression analysis (RT-PCR) using target-specific primers. The results obtained were compared with the in silico gene expression of the target genes in the TCGA datasets. Co-expression analysis was performed to establish an association between PTGS2 and VEGF signalling genes.
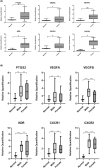
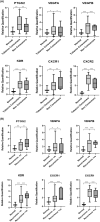
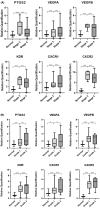
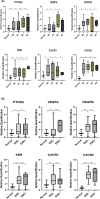
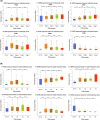
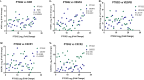
Results: Tumour and normal tissue samples were collected from 24 OSCC patients. Significant upregulation of PTGS2 expression was observed. Furthermore, VEGFA, KDR, CXCR1 and CXCR2 were significantly upregulated in tumour samples compared with paired normal samples, except for VEGFB, whose expression was not statistically significant. A similar expression pattern was observed in silico, except for CXCR2 which was highly expressed in the normal samples. Co-expression analysis showed a significant positive correlation between PTGS2 and VEGF signalling genes, except for VEGFB which showed a negative correlation.
Conclusion: PTGS2 and VEGF signalling genes are upregulated in OSCC, which has a profound impact on clinical outcomes.
Keywords: PTGS2; VEGF; Oral cancer; angiogenesis; gene expression; metastasis.
© 2024 The Authors. Cancer Medicine published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Prostaglandin-endoperoxide synthase 2 (PTGS2) gene expression and its association with genes regulating the VEGF signaling pathway in head and neck squamous cell carcinoma.J Oral Biol Craniofac Res. 2023 Sep-Oct;13(5):567-574. doi: 10.1016/j.jobcr.2023.07.002. Epub 2023 Jul 27. J Oral Biol Craniofac Res. 2023. PMID: 37559688 Free PMC article.
-
Role of hyaluronan mediated motility receptor gene in oral squamous cell carcinoma and clinical prognosis.Zhong Nan Da Xue Xue Bao Yi Xue Ban. 2021 Dec 28;46(12):1315-1324. doi: 10.11817/j.issn.1672-7347.2021.200955. Zhong Nan Da Xue Xue Bao Yi Xue Ban. 2021. PMID: 35232899 Free PMC article. Chinese, English.
-
Inverse correlation of CD3 & vascular endothelial growth factor in incisional oral squamous cell carcinoma biopsies predicts nodal metastasis & poor survival of patients.Indian J Med Res. 2023 May;157(5):438-446. doi: 10.4103/ijmr.IJMR_1560_19. Indian J Med Res. 2023. PMID: 37322630 Free PMC article.
-
HMG20A was identified as a key enhancer driver associated with DNA damage repair in oral squamous cell carcinomas.BMC Oral Health. 2022 Nov 5;22(1):473. doi: 10.1186/s12903-022-02500-y. BMC Oral Health. 2022. PMID: 36335317 Free PMC article.
-
Notch Signalling: A Potential Therapeutic Pathway in Oral Squamous Cell Carcinoma.Endocr Metab Immune Disord Drug Targets. 2021;21(12):2159-2168. doi: 10.2174/1871530321666210421125231. Endocr Metab Immune Disord Drug Targets. 2021. PMID: 33882814 Review.
Cited by
-
The key target and role of betel nut in oral squamous cell carcinoma.BMC Oral Health. 2025 Jul 19;25(1):1212. doi: 10.1186/s12903-025-06558-2. BMC Oral Health. 2025. PMID: 40684141 Free PMC article.
-
Physiological Effects of Soat1 Inactivation on Homeostasis of the Mouse Ocular Surface.Invest Ophthalmol Vis Sci. 2024 Jul 1;65(8):2. doi: 10.1167/iovs.65.8.2. Invest Ophthalmol Vis Sci. 2024. PMID: 38953847 Free PMC article.
-
Berries as Nature's Therapeutics: Exploring the Potential of Vaccinium Metabolites in Gastric Cancer Treatment Through Computational Insights.Life (Basel). 2025 Mar 5;15(3):406. doi: 10.3390/life15030406. Life (Basel). 2025. PMID: 40141751 Free PMC article.
-
Investigation of the Molecular Mechanisms of Paraoxonase-2 Mediated Radiotherapy and Chemotherapy Resistance in Oral Squamous Cell Carcinoma.Clin Transl Sci. 2025 Apr;18(4):e70201. doi: 10.1111/cts.70201. Clin Transl Sci. 2025. PMID: 40134131 Free PMC article. Review.
-
Antiapoptotic PON2 expression and its clinical implications in locally advanced oral squamous cell carcinoma.Cancer Sci. 2024 Jun;115(6):2012-2022. doi: 10.1111/cas.16170. Epub 2024 Apr 11. Cancer Sci. 2024. PMID: 38602182 Free PMC article.
References
-
- Mummudi N, Agarwal JP, Chatterjee S, Mallick I, Ghosh‐Laskar S. Oral cavity cancer in the Indian subcontinent – challenges and opportunities. Clin Oncol. 2019;31(8):520‐528. - PubMed
-
- Anil Kumar S, Indu S, Gautami D, Varruchi S. Oral squamous cell carcinoma (OSCC) in humans: etiological factors, diagnostic and therapeutic relevance. Res J Biotechnol. 2020;15:141‐151. https://www.researchgate.net/publication/344374174
-
- Patra S, Patil S, Das S, Bhutia SK. Epigenetic dysregulation in autophagy signaling as a driver of viral manifested oral carcinogenesis. Biochim Biophys Acta Mol basis Dis. 2022;1868(11):166517. - PubMed
-
- Al‐Ostoot FH, Salah S, Khamees HA, Khanum SA. Tumor angiogenesis: current challenges and therapeutic opportunities. Cancer Treat Res Commun. 2021;28:100422. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

