Follicular lymphoma B cells exhibit heterogeneous transcriptional states with associated somatic alterations and tumor microenvironments
- PMID: 38428430
- PMCID: PMC10983045
- DOI: 10.1016/j.xcrm.2024.101443
Follicular lymphoma B cells exhibit heterogeneous transcriptional states with associated somatic alterations and tumor microenvironments
Abstract
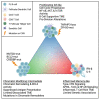
Follicular lymphoma (FL) is an indolent non-Hodgkin lymphoma of germinal center origin, which presents with significant biologic and clinical heterogeneity. Using RNA-seq on B cells sorted from 87 FL biopsies, combined with machine-learning approaches, we identify 3 transcriptional states that divide the biological ontology of FL B cells into inflamed, proliferative, and chromatin-modifying states, with relationship to prior GC B cell phenotypes. When integrated with whole-exome sequencing and immune profiling, we find that each state was associated with a combination of mutations in chromatin modifiers, copy-number alterations to TNFAIP3, and T follicular helper cells (Tfh) cell interactions, or primarily by a microenvironment rich in activated T cells. Altogether, these data define FL B cell transcriptional states across a large cohort of patients, contribute to our understanding of FL heterogeneity at the tumor cell level, and provide a foundation for guiding therapeutic intervention.
Keywords: B cell; follicular lymphoma; genomics; germinal center; transcriptome; tumor microenvironment.
Copyright © 2024 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests A.J.N. has received research funding from Bristol Myers Squibb.
Figures






References
-
- Roulland S., Faroudi M., Mamessier E., Sungalee S., Salles G., Nadel B. In: Advances in Immunology. Alt F.W., editor. Academic Press; 2011. Chapter 1 - Early Steps of Follicular Lymphoma Pathogenesis; pp. 1–46. - PubMed
-
- Cleary M.L., Sklar J. Nucleotide sequence of a t(14;18) chromosomal breakpoint in follicular lymphoma and demonstration of a breakpoint-cluster region near a transcriptionally active locus on chromosome 18. Proc. Natl. Acad. Sci. USA. 1985;82:7439–7443. doi: 10.1073/pnas.82.21.7439. - DOI - PMC - PubMed
-
- Nooka A.K., Nabhan C., Zhou X., Taylor M.D., Byrtek M., Miller T.P., Friedberg J.W., Zelenetz A.D., Link B.K., Cerhan J.R., et al. Examination of the follicular lymphoma international prognostic index (FLIPI) in the National LymphoCare study (NLCS): a prospective US patient cohort treated predominantly in community practices. Ann. Oncol. 2013;24:441–448. doi: 10.1093/annonc/mds429. - DOI - PubMed
-
- Dave S.S., Wright G., Tan B., Rosenwald A., Gascoyne R.D., Chan W.C., Fisher R.I., Braziel R.M., Rimsza L.M., Grogan T.M., et al. Prediction of Survival in Follicular Lymphoma Based on Molecular Features of Tumor-Infiltrating Immune Cells. N. Engl. J. Med. 2004;351:2159–2169. doi: 10.1056/nejmoa041869. - DOI - PubMed
-
- Huet S., Tesson B., Jais J.-P., Feldman A.L., Magnano L., Thomas E., Traverse-Glehen A., Albaud B., Carrère M., Xerri L., et al. A gene-expression profiling score for prediction of outcome in patients with follicular lymphoma: a retrospective training and validation analysis in three international cohorts. Lancet Oncol. 2018;19:549–561. doi: 10.1016/s1470-2045(18)30102-5. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

