Protein NirP1 regulates nitrite reductase and nitrite excretion in cyanobacteria
- PMID: 38429292
- PMCID: PMC10907346
- DOI: 10.1038/s41467-024-46253-4
Protein NirP1 regulates nitrite reductase and nitrite excretion in cyanobacteria
Abstract
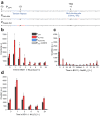
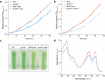
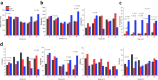
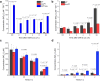
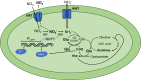
When the supply of inorganic carbon is limiting, photosynthetic cyanobacteria excrete nitrite, a toxic intermediate in the ammonia assimilation pathway from nitrate. It has been hypothesized that the excreted nitrite represents excess nitrogen that cannot be further assimilated due to the missing carbon, but the underlying molecular mechanisms are unclear. Here, we identified a protein that interacts with nitrite reductase, regulates nitrogen metabolism and promotes nitrite excretion. The protein, which we named NirP1, is encoded by an unannotated gene that is upregulated under low carbon conditions and controlled by transcription factor NtcA, a central regulator of nitrogen homeostasis. Ectopic overexpression of nirP1 in Synechocystis sp. PCC 6803 resulted in a chlorotic phenotype, delayed growth, severe changes in amino acid pools, and nitrite excretion. Coimmunoprecipitation experiments indicated that NirP1 interacts with nitrite reductase, a central enzyme in the assimilation of ammonia from nitrate/nitrite. Our results reveal that NirP1 is widely conserved in cyanobacteria and plays a crucial role in the coordination of C/N primary metabolism by targeting nitrite reductase.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
Nitrate/nitrite assimilation system of the marine picoplanktonic cyanobacterium Synechococcus sp. strain WH 8103: effect of nitrogen source and availability on gene expression.Appl Environ Microbiol. 2003 Dec;69(12):7009-18. doi: 10.1128/AEM.69.12.7009-7018.2003. Appl Environ Microbiol. 2003. PMID: 14660343 Free PMC article.
-
The roles of the nitrate reductase NarGHJI, the nitrite reductase NirBD and the response regulator GlnR in nitrate assimilation of Mycobacterium tuberculosis.Microbiology (Reading). 2009 Apr;155(Pt 4):1332-1339. doi: 10.1099/mic.0.023275-0. Microbiology (Reading). 2009. PMID: 19332834
-
Induction of the Nitrate Assimilation nirA Operon and Protein-Protein Interactions in the Maturation of Nitrate and Nitrite Reductases in the Cyanobacterium Anabaena sp. Strain PCC 7120.J Bacteriol. 2015 Jul;197(14):2442-52. doi: 10.1128/JB.00198-15. Epub 2015 May 11. J Bacteriol. 2015. PMID: 25962912 Free PMC article.
-
Nitrate assimilation by bacteria.Adv Microb Physiol. 1998;39:1-30, 379. doi: 10.1016/s0065-2911(08)60014-4. Adv Microb Physiol. 1998. PMID: 9328645 Review.
-
Photosynthetic nitrate assimilation in cyanobacteria.Photosynth Res. 2005;83(2):117-33. doi: 10.1007/s11120-004-5830-9. Photosynth Res. 2005. PMID: 16143847 Review.
Cited by
-
Epigenetic control of tetrapyrrole biosynthesis by m4C DNA methylation in a cyanobacterium.DNA Res. 2024 Dec 1;31(6):dsae035. doi: 10.1093/dnares/dsae035. DNA Res. 2024. PMID: 39657587 Free PMC article.
-
The primary carbon metabolism in cyanobacteria and its regulation.Front Plant Sci. 2024 Jul 5;15:1417680. doi: 10.3389/fpls.2024.1417680. eCollection 2024. Front Plant Sci. 2024. PMID: 39036361 Free PMC article. Review.
-
Electron transfer in biological systems.J Biol Inorg Chem. 2024 Dec;29(7-8):641-683. doi: 10.1007/s00775-024-02076-8. Epub 2024 Oct 18. J Biol Inorg Chem. 2024. PMID: 39424709 Free PMC article. Review.
-
How Small Proteins Adjust the Metabolism of Cyanobacteria Under Stress: The Role of Small Proteins in Cyanobacterial Stress Responses.Bioessays. 2025 Mar;47(3):e202400245. doi: 10.1002/bies.202400245. Epub 2024 Dec 12. Bioessays. 2025. PMID: 39668401 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

