To initiate or not to initiate: A critical assessment of eIF2A, eIF2D, and MCT-1·DENR to deliver initiator tRNA to ribosomes
- PMID: 38433101
- PMCID: PMC11260288
- DOI: 10.1002/wrna.1833
To initiate or not to initiate: A critical assessment of eIF2A, eIF2D, and MCT-1·DENR to deliver initiator tRNA to ribosomes
Abstract
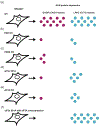
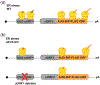
Selection of the correct start codon is critical for high-fidelity protein synthesis. In eukaryotes, this is typically governed by a multitude of initiation factors (eIFs), including eIF2·GTP that directly delivers the initiator tRNA (Met-tRNAi Met ) to the P site of the ribosome. However, numerous reports, some dating back to the early 1970s, have described other initiation factors having high affinity for the initiator tRNA and the ability of delivering it to the ribosome, which has provided a foundation for further work demonstrating non-canonical initiation mechanisms using alternative initiation factors. Here we provide a critical analysis of current understanding of eIF2A, eIF2D, and the MCT-1·DENR dimer, the evidence surrounding their ability to initiate translation, their implications in human disease, and lay out important key questions for the field. This article is categorized under: RNA Interactions with Proteins and Other Molecules > RNA-Protein Complexes Translation > Mechanisms Translation > Regulation.
Keywords: eIF; translation initiation; translational control.
© 2024 The Authors. WIREs RNA published by Wiley Periodicals LLC.
Conflict of interest statement
CONFLICT OF INTEREST STATEMENT
The authors declare no conflicts of interest.
Figures


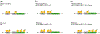
Similar articles
-
Structural and Functional Insights into Human Re-initiation Complexes.Mol Cell. 2017 Aug 3;67(3):447-456.e7. doi: 10.1016/j.molcel.2017.06.032. Epub 2017 Jul 18. Mol Cell. 2017. PMID: 28732596
-
Activities of Ligatin and MCT-1/DENR in eukaryotic translation initiation and ribosomal recycling.Genes Dev. 2010 Aug 15;24(16):1787-801. doi: 10.1101/gad.1957510. Genes Dev. 2010. PMID: 20713520 Free PMC article.
-
Hepatitis C Virus Translation Regulation.Int J Mol Sci. 2020 Mar 27;21(7):2328. doi: 10.3390/ijms21072328. Int J Mol Sci. 2020. PMID: 32230899 Free PMC article. Review.
-
eIF2A, an initiator tRNA carrier refractory to eIF2α kinases, functions synergistically with eIF5B.Cell Mol Life Sci. 2018 Dec;75(23):4287-4300. doi: 10.1007/s00018-018-2870-4. Epub 2018 Jul 17. Cell Mol Life Sci. 2018. PMID: 30019215 Free PMC article.
-
Structural insights into eukaryotic ribosomes and the initiation of translation.Curr Opin Struct Biol. 2012 Dec;22(6):768-77. doi: 10.1016/j.sbi.2012.07.010. Epub 2012 Aug 10. Curr Opin Struct Biol. 2012. PMID: 22889726 Review.
Cited by
-
MCTS2 and distinct eIF2D roles in uORF-dependent translation regulation revealed by in vitro re-initiation assays.EMBO J. 2025 Feb;44(3):854-876. doi: 10.1038/s44318-024-00347-3. Epub 2025 Jan 2. EMBO J. 2025. PMID: 39748120 Free PMC article.
-
Nonstop mutations cause loss of renal tumor suppressor proteins VHL and BAP1 and affect multiple stages of protein translation.Sci Adv. 2025 Feb 14;11(7):eadr6375. doi: 10.1126/sciadv.adr6375. Epub 2025 Feb 12. Sci Adv. 2025. PMID: 39937911 Free PMC article.
-
lncRNAs: the unexpected link between protein synthesis and cancer adaptation.Mol Cancer. 2025 Jan 31;24(1):38. doi: 10.1186/s12943-025-02236-7. Mol Cancer. 2025. PMID: 39891197 Free PMC article. Review.
-
Characterizing differences in the muscle transcriptome between cattle with alternative LCORL-NCAPG haplotypes.BMC Genomics. 2025 May 14;26(1):479. doi: 10.1186/s12864-025-11665-z. BMC Genomics. 2025. PMID: 40369436 Free PMC article.
-
Human eIF2A has a minimal role in translation initiation and in uORF-mediated translational control in HeLa cells.Elife. 2025 Jul 2;14:RP105311. doi: 10.7554/eLife.105311. Elife. 2025. PMID: 40600802 Free PMC article.
References
-
- Adams SL, Safer B, Anderson WF, & Merrick WC (1975). Eukaryotic initiation complex formation. Evidence for two distinct pathways. The Journal of Biological Chemistry, 250(23), 9083–9089. - PubMed
-
- Anderson R, Agarwal A, Ghosh A, Guan BJ, Casteel J, Dvorina N, Baldwin WM, Mazumder B, Nazarko TY, Merrick WC, Buchner DA, Hatzoglou M, Kondratov RV, & Komar AA (2021). eIF2A-knockout mice reveal decreased life span and metabolic syndrome. The FASEB Journal, 35(11), e21990. 10.1096/fj.202101105R - DOI - PMC - PubMed
-
- Arnaud E, Touriol C, Boutonnet C, Gensac MC, Vagner S, Prats H, & Prats AC (1999). A new 34-kilodalton isoform of human fibroblast growth factor 2 is cap dependently synthesized by using a non-AUG start codon and behaves as a survival factor. Molecular and Cellular Biology, 19(1), 505–514. 10.1128/MCB.19.1.505 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

