Regulation mechanisms of disulfidptosis-related genes in ankylosing spondylitis and inflammatory bowel disease
- PMID: 38433839
- PMCID: PMC10904683
- DOI: 10.3389/fimmu.2024.1326354
Regulation mechanisms of disulfidptosis-related genes in ankylosing spondylitis and inflammatory bowel disease
Abstract
Introduction: Disulfidptosis is a recently identified form of cell death that contributes to maintaining the internal environment balance of an organism. However, the molecular basis of disulfidptosis in ulcerative colitis (UC), ankylosing spondylitis (AS), and Crohn's disease (CD) has not been thoroughly explored.
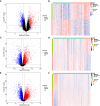
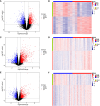
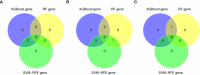
Methods: Firstly, the differentially expressed genes (DEGs) and disulfidptosis-associated genes (DAGs) were obtained through differential analysis between diseases (AS, CD, and UC) and control groups. After the disulfidptosis score was acquired using the single-sample gene set enrichment analysis (ssGSEA) algorithm, the DE-DAGs were screened by overlapping DAGs and DEGs of the three diseases. Next, the feature genes were selected through a combination of machine learning algorithms, receiver operating characteristic (ROC) curves, and expression analysis. Based on these feature genes, nomograms were created for AS, CD and UC. The co-feature genes were then identified by taking the intersections of the genes featured in all three diseases. Meanwhile, single-gene set enrichment analysis (GSEA) and the TF-mRNA-miRNA network were utilized to investigate the molecular mechanisms of the co-feature genes. To validate the expression differences of the co-feature genes between healthy controls and patients (AS and IBD), RT-PCR was performed. Lastly, mendelian randomization (MR) analysis was utilized to explore the causality between genetic variants of S100A12 with AS, UC and CD.
Results: In this study, 11 DE-DAGs were obtained. Functional enrichment analysis revealed their involvement in cytokine production and fatty acid biosynthesis. Latterly, AS/CD/UC -feature genes were derived, and they all had decent diagnostic performance. Through evaluation, the performance of the nomogram was decent for three diseases. Then, 2 co-feature genes (S100A12 and LILRA5) were obtained. The GSEA enrichment results indicated that the co-feature genes were mainly enriched in the cytokine-cytokine receptor interaction and drug metabolism cytochrome P450. As shown by functional experiments, there was a correlation between the mRNA expression of S100A12 with AS, UC and CD. Additionally, a causal connection between S100A12 and IBD was detected through MR analysis.
Discussion: In this study, 2 co-feature genes (S100A12 and LILRA5) were screened, and their functions were investigated in AS, CD and UC, providing a basis for further research into diagnosis and treatment.
Keywords: Crohn’s disease; ankylosing spondylitis; bioinformatics; disulfidptosis; mendelian randomization; ulcerative colitis.
Copyright © 2024 Li, Fang, Li, Xie, Zhou, Zhu, Jin, Song, Yang and Liping.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures











References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

