Analysis of m6A-related lncRNAs for prognostic and immunotherapeutic response in hepatocellular carcinoma
- PMID: 38434979
- PMCID: PMC10905389
- DOI: 10.7150/jca.92128
Analysis of m6A-related lncRNAs for prognostic and immunotherapeutic response in hepatocellular carcinoma
Abstract
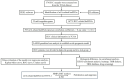
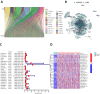
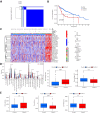
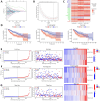
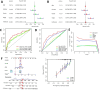
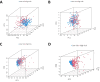
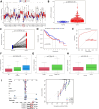
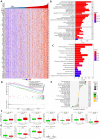
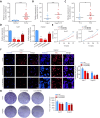
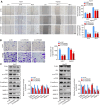
Background: RNA methylation modifications are important post-translational modifications that are regulated in an epigenetic manner. Recently, N6-methyladenosine (m6A) RNA modifications have emerged as potential epigenetic markers in tumor biology. Methods: Gene expression and clinicopathological data of LIHC were obtained from the cancer genome atlas (TCGA) database. The relationship between long non-coding RNAs (lncRNAs) and m6A-related genes was determined by gene expression analysis using Perl and R software. Co-expression network of m6A-lncRNA was constructed, and the relevant lncRNAs associated with prognosis were identified using univariate Cox regression analysis. These lncRNAs were then divided into two clusters (cluster 1 and cluster 2) to determine the differences in survival, pathoclinical parameters, and immune cell infiltration between the different lncRNA subtypes. The least absolute shrinkage and selection operator (LASSO) was carried out for regression analysis and prognostic model. The HCC patients were randomly divided into a train group and a test group. According to the median risk score of the model, HCC patients were divided into high-risk and low-risk groups. We built models using the train group and confirmed them through the test group. The m6A-lncRNAs derived from the models were analyzed for the tumor mutational burden (TMB), immune evasion and immune function using R software. AL355574.1 was identified as an important m6A-associated lncRNA and selected for further investigation. Finally, in vitro experiments were conducted to confirm the effect of AL355574.1 on the biological function of HCC and the possible biological mechanisms. Huh7 and HepG2 cells were transfected with AL355574.1 siRNA and cell proliferation ability was measured by CCK-8, EdU and colony formation assays. Wound healing and transwell assays were used to determine the cell migration capacity. The expression levels of MMP-2, MMP-9, E-cadherin, N-cadherin and Akt/mTOR phosphorylation were all determined by Western blotting. Results: The lncRNAs with significant prognostic value were classified into two subtypes by a consistent clustering analysis. We found that the clinical features, immune cell infiltration and tumor microenvironment (TME) were significantly different between the lncRNA subtypes. Our analysis revealed significant correlations between these different lncRNA subtypes and immune infiltrating and stromal cells. We created the final risk profile using LASSO regression, which notably included three lncRNAs (AL355574.1, AL158166.1, TMCC1-AS1). A prognostic signature consisting of the three lncRNAs was constructed, and the model showed excellent prognostic predictive ability. The overall survival (OS) of the low-risk cohort was significantly higher than that of the high-risk cohort in both the train and test group. Both risk score [hazard ratio (HR)=1.062; P<0.001] and stage (HR=1.647; P< 0.001) were considered independent indicators of HCC prognosis by univariate and multivariate Cox regression analysis. In Huh7 and HepG2 cells, AL355574.1 knockdown inhibited cell proliferation and migration, suppressed the protein expression levels of MMP-2, MMP-9, N-cadherin and Akt/mTOR phosphorylation, but promoted the protein expression levels of E-cadherin. Conclusions: This study established a predictive model for the OS of HCC patients, and these OS-related m6A-lncRNAs, especially AL355574.1 may play a potential role in the progression of HCC. In vitro experiments also showed that AL355574.1 could enhance the expression of MMPs and EMT through the Akt/mTOR signaling pathway, thereby affected the proliferation and migration of HCC. This provides a new perspective on the anticancer molecular mechanism of AL355574.1 in HCC.
Keywords: HCC; LASSO; immune; lncRNAs; m6A.
© The author(s).
Conflict of interest statement
Competing Interests: The authors have declared that no competing interest exists.
Figures

Similar articles
-
A Cuproptosis-Related LncRNA Risk Model for Predicting Prognosis and Immunotherapeutic Efficacy in Patients with Hepatocellular Carcinoma.Biochem Genet. 2024 Jun;62(3):2332-2351. doi: 10.1007/s10528-023-10539-x. Epub 2023 Oct 29. Biochem Genet. 2024. PMID: 37898914
-
Systematic analysis of the role of LDHs subtype in pan-cancer demonstrates the importance of LDHD in the prognosis of hepatocellular carcinoma patients.BMC Cancer. 2024 Jan 31;24(1):156. doi: 10.1186/s12885-024-11920-8. BMC Cancer. 2024. PMID: 38291366 Free PMC article.
-
Prediction of prognosis, immune infiltration, and personalized treatment of hepatocellular carcinoma by analysis of cuproptosis-related long noncoding RNAs and verification in vitro.Front Oncol. 2023 Sep 8;13:1159126. doi: 10.3389/fonc.2023.1159126. eCollection 2023. Front Oncol. 2023. PMID: 37746284 Free PMC article.
-
Function of the Long Noncoding RNAs in Hepatocellular Carcinoma: Classification, Molecular Mechanisms, and Significant Therapeutic Potentials.Bioengineering (Basel). 2022 Aug 21;9(8):406. doi: 10.3390/bioengineering9080406. Bioengineering (Basel). 2022. PMID: 36004931 Free PMC article. Review.
-
The regulation of hypoxia-related lncRNAs in hepatocellular carcinoma.Discov Oncol. 2024 May 7;15(1):144. doi: 10.1007/s12672-024-01002-3. Discov Oncol. 2024. PMID: 38713276 Free PMC article. Review.
Cited by
-
Identification and validation of a five-necroptosis-related lncRNAs signature for prognostic prediction in hepatocellular carcinoma.Heliyon. 2024 Sep 6;10(18):e37403. doi: 10.1016/j.heliyon.2024.e37403. eCollection 2024 Sep 30. Heliyon. 2024. PMID: 39309864 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

