Construction of a broad-host-range Anderson promoter series and particulate methane monooxygenase promoter variants expand the methanotroph genetic toolbox
- PMID: 38435708
- PMCID: PMC10909576
- DOI: 10.1016/j.synbio.2024.02.003
Construction of a broad-host-range Anderson promoter series and particulate methane monooxygenase promoter variants expand the methanotroph genetic toolbox
Abstract
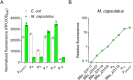
Methanotrophic bacteria are currently used industrially for the bioconversion of methane-rich natural gas and anaerobic digestion-derived biogas to valuable products. These bacteria may also serve to mitigate the negative effects of climate change by capturing atmospheric greenhouse gases. Several genetic tools have previously been developed for genetic and metabolic engineering of methanotrophs. However, the available tools for use in methanotrophs are significantly underdeveloped compared to many other industrially relevant bacteria, which hinders genetic and metabolic engineering of these biocatalysts. As such, expansion of the methanotroph genetic toolbox is needed to further our understanding of methanotrophy and develop biotechnologies that leverage these unique microbes for mitigation and conversion of methane to valuable products. Here, we determined the copy number of three broad-host-range plasmids in Methylococcus capsulatus Bath and Methylosinus trichosporium OB3b, representing phylogenetically diverse Gammaproteobacterial and Alphaproteobacterial methanotrophs, respectively. Further, we show that the commonly used synthetic Anderson series promoters are functional and exhibit similar relative activity in M. capsulatus and M. trichosporium OB3b, but the synthetic series had limited range. Thus, we mutagenized the native M. capsulatus particulate methane monooxygenase promoter and identified variants with activity that expand the activity range of synthetic, constitutive promoters functional not only in M. capsulatus, but also in Escherichia coli. Collectively, the tools developed here advance the methanotroph genetic engineering toolbox and represent additional synthetic genetic parts that may have broad applicability in Pseudomonadota bacteria.
Keywords: Metabolic engineering; Methane monooxygenase; Methanotroph; Promoter; Synthetic biology.
© 2024 The Authors.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures




Similar articles
-
Development of a CRISPR/Cas9 System for Methylococcus capsulatus In Vivo Gene Editing.Appl Environ Microbiol. 2019 May 16;85(11):e00340-19. doi: 10.1128/AEM.00340-19. Print 2019 Jun 1. Appl Environ Microbiol. 2019. PMID: 30926729 Free PMC article.
-
Methane Monooxygenase Gene Transcripts as Quantitative Biomarkers of Methanotrophic Activity in Methylosinus trichosporium OB3b.Appl Environ Microbiol. 2020 Nov 10;86(23):e01048-20. doi: 10.1128/AEM.01048-20. Print 2020 Nov 10. Appl Environ Microbiol. 2020. PMID: 32948519 Free PMC article.
-
Molecular analysis of the pmo (particulate methane monooxygenase) operons from two type II methanotrophs.Appl Environ Microbiol. 2000 Mar;66(3):966-75. doi: 10.1128/AEM.66.3.966-975.2000. Appl Environ Microbiol. 2000. PMID: 10698759 Free PMC article.
-
Methanotrophs, Methylosinus trichosporium OB3b, sMMO, and their application to bioremediation.Crit Rev Microbiol. 1998;24(4):335-73. doi: 10.1080/10408419891294217. Crit Rev Microbiol. 1998. PMID: 9887367 Review.
-
Recent findings in methanotrophs: genetics, molecular ecology, and biopotential.Appl Microbiol Biotechnol. 2024 Dec;108(1):60. doi: 10.1007/s00253-023-12978-3. Epub 2024 Jan 6. Appl Microbiol Biotechnol. 2024. PMID: 38183483 Review.
Cited by
-
Construction of a synthetic biology toolkit for the genetic manipulation in the cellulose-producing strain Kosakonia oryzendophytica.Synth Syst Biotechnol. 2025 May 27;10(3):1050-1058. doi: 10.1016/j.synbio.2025.05.012. eCollection 2025 Sep. Synth Syst Biotechnol. 2025. PMID: 40529626 Free PMC article.
-
A genetically encoded biosensor to monitor dynamic changes of c-di-GMP with high temporal resolution.Nat Commun. 2024 May 9;15(1):3920. doi: 10.1038/s41467-024-48295-0. Nat Commun. 2024. PMID: 38724508 Free PMC article.
References
-
- Nariya S., Kalyuzhnaya M.G. Hemerythrins enhance aerobic respiration in Methylomicrobium alcaliphilum 20ZR, a methane-consuming bacterium. FEMS Microbiol Lett. 2020;367 - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

