Identification of plants' functional counterpart of the metazoan mediator of DNA Damage checkpoint 1
- PMID: 38438802
- PMCID: PMC11014961
- DOI: 10.1038/s44319-024-00107-8
Identification of plants' functional counterpart of the metazoan mediator of DNA Damage checkpoint 1
Abstract
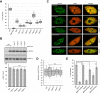
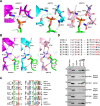
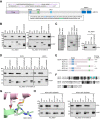
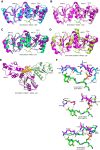
Induction of DNA damage triggers rapid phosphorylation of the histone H2A.X (γH2A.X). In animals, mediator of DNA damage checkpoint 1 (MDC1) binds γH2A.X through a tandem BRCA1 carboxyl-terminal (tBRCT) domain and mediates recruitment of downstream effectors of DNA damage response (DDR). However, readers of this modification in plants have remained elusive. We show that from the Arabidopsis BRCT domain proteome, BCP1-4 proteins with tBRCT domains are involved in DDR. Through its tBRCT domain BCP4 binds γH2A.X in vitro and localizes to DNA damage-induced foci in an H2A.X-dependent manner. BCP4 also contains a domain that interacts directly with NBS1 and thus acts as a functional counterpart of MDC1. We also show that BCP1, that contains two tBRCT domains, co-localizes with γH2A.X but it does not bind γH2A.X suggesting functional similarity with human PAXIP1. A phylogenetic analysis supports that PAXIP1 and MDC1 in metazoa and their plant counterparts evolved independently from common ancestors with tBRCT domains. Collectively, our study reveals missing components and provides mechanistic and evolutionary insights into plant DDR.
Keywords: BCP4; BRCT Domain; DNA Damage Response; Histone H2A.X; MDC1.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures







Similar articles
-
Mdb1, a fission yeast homolog of human MDC1, modulates DNA damage response and mitotic spindle function.PLoS One. 2014 May 7;9(5):e97028. doi: 10.1371/journal.pone.0097028. eCollection 2014. PLoS One. 2014. PMID: 24806815 Free PMC article.
-
Structure of a second BRCT domain identified in the nijmegen breakage syndrome protein Nbs1 and its function in an MDC1-dependent localization of Nbs1 to DNA damage sites.J Mol Biol. 2008 Aug 29;381(2):361-72. doi: 10.1016/j.jmb.2008.05.087. Epub 2008 Jun 14. J Mol Biol. 2008. PMID: 18582474 Free PMC article.
-
MDC1 is a mediator of the mammalian DNA damage checkpoint.Nature. 2003 Feb 27;421(6926):961-6. doi: 10.1038/nature01446. Nature. 2003. PMID: 12607005
-
Molecular mechanism of the recruitment of NBS1/hMRE11/hRAD50 complex to DNA double-strand breaks: NBS1 binds to gamma-H2AX through FHA/BRCT domain.J Radiat Res. 2004 Dec;45(4):473-8. doi: 10.1269/jrr.45.473. J Radiat Res. 2004. PMID: 15635255 Review.
-
MDC1/NFBD1: a key regulator of the DNA damage response in higher eukaryotes.DNA Repair (Amst). 2004 Aug-Sep;3(8-9):953-7. doi: 10.1016/j.dnarep.2004.03.007. DNA Repair (Amst). 2004. PMID: 15279781 Review.
Cited by
-
Chromatin dynamics and RNA metabolism are double-edged swords for the maintenance of plant genome integrity.Nat Plants. 2024 Jun;10(6):857-873. doi: 10.1038/s41477-024-01678-z. Epub 2024 Apr 24. Nat Plants. 2024. PMID: 38658791 Review.
-
The PST repeat region of MDC1 is a tunable multivalent chromatin tethering domain.bioRxiv [Preprint]. 2025 Jan 13:2025.01.10.632395. doi: 10.1101/2025.01.10.632395. bioRxiv. 2025. PMID: 39868091 Free PMC article. Preprint.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

