Optimal control of ribosome population for gene expression under periodic nutrient intake
- PMID: 38442858
- PMCID: PMC10914516
- DOI: 10.1098/rsif.2023.0652
Optimal control of ribosome population for gene expression under periodic nutrient intake
Abstract
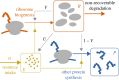
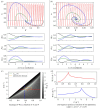
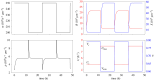
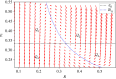
Translation of proteins is a fundamental part of gene expression that is mediated by ribosomes. As ribosomes significantly contribute to both cellular mass and energy consumption, achieving efficient management of the ribosome population is also crucial to metabolism and growth. Inspired by biological evidence for nutrient-dependent mechanisms that control both ribosome-active degradation and genesis, we introduce a dynamical model of protein production, that includes the dynamics of resources and control over the ribosome population. Under the hypothesis that active degradation and biogenesis are optimal for maximizing and maintaining protein production, we aim to qualitatively reproduce empirical observations of the ribosome population dynamics. Upon formulating the associated optimization problem, we first analytically study the stability and global behaviour of solutions under constant resource input, and characterize the extent of oscillations and convergence rate to a global equilibrium. We further use these results to simplify and solve the problem under a quasi-static approximation. Using biophysical parameter values, we find that optimal control solutions lead to both control mechanisms and the ribosome population switching between periods of feeding and fasting, suggesting that the intense regulation of ribosome population observed in experiments allows to maximize and maintain protein production. Finally, we find some range for the control values over which such a regime can be observed, depending on the intensity of fasting.
Keywords: dynamical system; optimal control; protein translation; ribophagy; ribosome; systems biology.
Conflict of interest statement
This work did not require ethical approval from a human subject or animal welfare committee.
We declare we have no competing interests.
Figures

Similar articles
-
Ribosome flow model with extended objects.J R Soc Interface. 2017 Oct;14(135):20170128. doi: 10.1098/rsif.2017.0128. J R Soc Interface. 2017. PMID: 29021157 Free PMC article.
-
Maximizing protein translation rate in the non-homogeneous ribosome flow model: a convex optimization approach.J R Soc Interface. 2014 Nov 6;11(100):20140713. doi: 10.1098/rsif.2014.0713. J R Soc Interface. 2014. PMID: 25232050 Free PMC article.
-
Maximizing Protein Translation Rate in the Ribosome Flow Model: The Homogeneous Case.IEEE/ACM Trans Comput Biol Bioinform. 2014 Nov-Dec;11(6):1184-95. doi: 10.1109/TCBB.2014.2330621. IEEE/ACM Trans Comput Biol Bioinform. 2014. PMID: 26357054
-
Eukaryotic ribosome quality control system: a potential therapeutic target for human diseases.Int J Biol Sci. 2022 Mar 14;18(6):2497-2514. doi: 10.7150/ijbs.70955. eCollection 2022. Int J Biol Sci. 2022. PMID: 35414791 Free PMC article. Review.
-
Ribosome dynamics and mRNA turnover, a complex relationship under constant cellular scrutiny.Wiley Interdiscip Rev RNA. 2021 Nov;12(6):e1658. doi: 10.1002/wrna.1658. Epub 2021 May 5. Wiley Interdiscip Rev RNA. 2021. PMID: 33949788 Free PMC article. Review.
Cited by
-
Dynamic optimization elucidates higher-level pathogenicity strategies of Pseudomonas aeruginosa.Microlife. 2025 Mar 13;6:uqaf005. doi: 10.1093/femsml/uqaf005. eCollection 2025. Microlife. 2025. PMID: 40182079 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials

