Morphological entropy encodes cellular migration strategies on multiple length scales
- PMID: 38453929
- PMCID: PMC10920856
- DOI: 10.1038/s41540-024-00353-5
Morphological entropy encodes cellular migration strategies on multiple length scales
Abstract
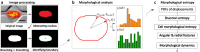
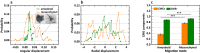
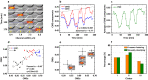
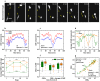
Cell migration is crucial for numerous physiological and pathological processes. A cell adapts its morphology, including the overall and nuclear morphology, in response to various cues in complex microenvironments, such as topotaxis and chemotaxis during migration. Thus, the dynamics of cellular morphology can encode migration strategies, from which diverse migration mechanisms can be inferred. However, deciphering the mechanisms behind cell migration encoded in morphology dynamics remains a challenging problem. Here, we present a powerful universal metric, the Cell Morphological Entropy (CME), developed by combining parametric morphological analysis with Shannon entropy. The utility of CME, which accurately quantifies the complex cellular morphology at multiple length scales through the deviation from a perfectly circular shape, is illustrated using a variety of normal and tumor cell lines in different in vitro microenvironments. Our results show how geometric constraints affect the MDA-MB-231 cell nucleus, the emerging interactions of MCF-10A cells migrating on collagen gel, and the critical transition from proliferation to invasion in tumor spheroids. The analysis demonstrates that the CME-based approach provides an effective and physically interpretable tool to measure morphology in real-time across multiple length scales. It provides deeper insight into cell migration and contributes to the understanding of different behavioral modes and collective cell motility in more complex microenvironments.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
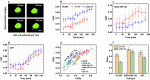
References
MeSH terms
Grants and funding
- 12347178/National Natural Science Foundation of China (National Science Foundation of China)
- 62106032/National Natural Science Foundation of China (National Science Foundation of China)
- 12090052/National Natural Science Foundation of China (National Science Foundation of China)
- 62171073/National Natural Science Foundation of China (National Science Foundation of China)
- CSTB2022NSCQ-MSX1260/Natural Science Foundation of Chongqing (Natural Science Foundation of Chongqing Municipality)
LinkOut - more resources
Full Text Sources
Miscellaneous

