Genetic relationships between sweet cherry (Prunus avium L.) and sour cherry (P. cerasus L.) as revealed using fruit characterizations and chloroplast microsatellites
- PMID: 38455164
- PMCID: PMC10916583
- DOI: 10.1002/fsn3.3858
Genetic relationships between sweet cherry (Prunus avium L.) and sour cherry (P. cerasus L.) as revealed using fruit characterizations and chloroplast microsatellites
Abstract
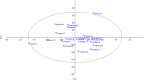
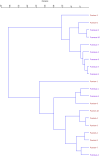
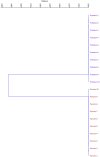
In the present study, the genetic diversity as well as the relationship between sweet cherry (Prunus avium L.) and sour cherry (P. cerasus L.) genotypes were investigated based on fruit traits and chloroplast microsatellites (cpSSRs). Analysis of variance showed that the studied genotypes have significant differences in the studied traits. In sweet cherries, the average fruit weight was 4.49 g with a coefficient of variation (CV) of 15.62%, the average stone weight was 0.34 g with a CV of 15.67%, and the average total soluble solids was 11.90% with a CV of 22.06%. Also, in sour cherries, the average fruit weight was 2.65 g with a coefficient of variation (CV) of 14.27%, the average stone weight was 0.28 g with a CV of 12.27%, and the average total soluble solids was 10.90% with a CV of 19.80%. Principal component analysis (PCA) showed that 83.80% of the observed variance was explained by the first three components. The cluster analysis separated genotypes of sweet and sour cherries and put them into two main groups. Four cpSSR primers produced distinct and different alleles among sweet and sour cherries. The cpSSR loci separated sweet and sour cherries from each other, which confirms the theory that chloroplast genome of sour cherry is not derived from sweet cherry. The present results provided new insights regarding the extent of diversity of individuals and also determined the relatedness and obtained information on genetic diversity of sweet and sour cherries.
Keywords: breeding; cherries; chloroplast; fruit; variation.
© 2023 The Authors. Food Science & Nutrition published by Wiley Periodicals LLC.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
Similar articles
-
Investigation of pollen morphology and viability of sweet and sour cherry genotypes by multivariate analysis.Microsc Res Tech. 2025 Jan;88(1):42-52. doi: 10.1002/jemt.24674. Epub 2024 Aug 23. Microsc Res Tech. 2025. PMID: 39177054
-
Chloroplast inheritance and DNA variation in sweet, sour, and ground cherry.J Hered. 2000 Jan-Feb;91(1):75-9. doi: 10.1093/jhered/91.1.75. J Hered. 2000. PMID: 10739133
-
Identification of incompatibility alleles in the tetraploid species sour cherry.Theor Appl Genet. 2004 Mar;108(5):775-85. doi: 10.1007/s00122-003-1511-x. Epub 2003 Dec 19. Theor Appl Genet. 2004. PMID: 14689184
-
Methods for quality evaluation of sweet cherry.J Sci Food Agric. 2023 Jan 30;103(2):463-478. doi: 10.1002/jsfa.12144. Epub 2022 Aug 15. J Sci Food Agric. 2023. PMID: 35870155 Review.
-
From Orchard to Wellness: Unveiling the Health Effects of Sweet Cherry Nutrients.Nutrients. 2024 Oct 28;16(21):3660. doi: 10.3390/nu16213660. Nutrients. 2024. PMID: 39519493 Free PMC article. Review.
Cited by
-
De novo assembly and comparative analysis of cherry (Prunus subgenus Cerasus) mitogenomes.Front Plant Sci. 2025 Mar 24;16:1568698. doi: 10.3389/fpls.2025.1568698. eCollection 2025. Front Plant Sci. 2025. PMID: 40196431 Free PMC article.
References
-
- Ates, U. , & Ozturk, B. (2022). Fruit quality characteristics of different sweet cherry (Prunus avium L.) cultivars grown in Ordu province of Turkey. Karadeniz Fen Bilimleri Dergisi, 12(1), 168–177.
-
- Brettin, T. S. , Karle, R. , Crowe, E. L. , & Iezzoni, F. (2000). Chloroplast inheritance and DNA variation in sweet, sour and ground cherry. Heredity, 91, 74–79. - PubMed
-
- Brown, A. H. D. (1978). Isozymes, plant population genetic structure and genetic conservation. Theoretical and Applied Genetics, 52, 145–157. - PubMed
-
- Decroocq, V. , Hagen, L. S. , Favé, M. G. , Eyquard, J. P. , & Pierronnet, A. (2004). Microsatellite markers in the hexaploid Prunus domestica species and parentage lineage of three European plum cultivars using nuclear and chloroplast simple‐sequence repeats. Molecular Breeding, 13, 135–142.
-
- Doyle, J. J. , & Doyle, J. L. (1987). Isolation of DNA from fresh plant tissue. Focus, 12, 13–15.
LinkOut - more resources
Full Text Sources