Stroke Causes DNA Methylation at Ncx1 Heart Promoter in the Brain Via DNMT1/MeCP2/REST Epigenetic Complex
- PMID: 38456444
- PMCID: PMC11010005
- DOI: 10.1161/JAHA.123.030460
Stroke Causes DNA Methylation at Ncx1 Heart Promoter in the Brain Via DNMT1/MeCP2/REST Epigenetic Complex
Abstract
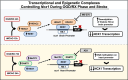
Background: REST (Repressor-Element 1 [RE1]-silencing transcription factor) inhibits Na+/Ca2+exchanger-1 (Ncx1) transcription in neurons through the binding of RE1 site on brain promoter (Br) after stroke. We identified a new putative RE1 site in Ncx1 heart promoter (Ht) sequence (Ht-RE1) that participates in neuronal Ncx1 transcription. Because REST recruits DNA-methyltransferase-1 (DNMT1) and MeCP2 (methyl-CpG binding protein 2) on different neuronal genes, we investigated the role of this complex in Ncx1 transcriptional regulation after stroke.
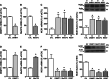
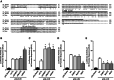
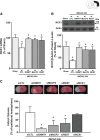
Methods and results: Luciferase experiments performed in SH-SY5Y cells demonstrated that Br activity was selectively decreased by REST, whereas Ht activity was reduced by DNMT1, MeCP2, and REST. Notably, site-direct mutagenesis of Ht-RE1 prevented REST-dependent downregulation of Ncx1. Furthermore, in temporoparietal cortex of 8-week-old male wild-type mice (C57BL/6) subjected to transient middle cerebral artery occlusion, DNMT1, MeCP2, and REST binding to Ht promoter was increased, with a consequent DNA promoter hypermethylation. Intracerebroventricular injection of siREST prevented DNMT1/MeCP2 binding to Ht and Ncx1 downregulation, thus causing a reduction in stroke-induced damage. Consistently, in cortical neurons subjected to oxygen and glucose deprivation plus reoxygenation Ncx1 knockdown counteracted neuronal protection induced by the demethylating agent 5-azacytidine. For comparisons between 2 experimental groups, Student's t test was used, whereas for more than 2 experimental groups, 1-way ANOVA was used, followed by Tukey or Newman Keuls. Statistical significance was set at P<0.05.
Conclusions: If the results of this study are confirmed in humans, it could be asserted that DNMT1/MeCP2/REST complex disruption could be a new pharmacological strategy to reduce DNA methylation of Ht in the brain, ameliorating stroke damage.
Keywords: DNA methylation; DNMT1; MeCP2; NCX1; REST; stroke.
Figures



References
-
- Molinaro P, Sirabella R, Pignataro G, Petrozziello T, Secondo A, Boscia F, Vinciguerra A, Cuomo O, Philipson KD, De Felice M, et al. Neuronal NCX1 overexpression induces stroke resistance while knockout induces vulnerability via Akt. J Cereb Blood Flow Metab. 2016;36:1790–1803. doi: 10.1177/0271678X15611913 - DOI - PMC - PubMed
-
- Formisano L, Guida N, Valsecchi V, Cantile M, Cuomo O, Vinciguerra A, Laudati G, Pignataro G, Sirabella R, Di Renzo G, et al. Sp3/REST/HDAC1/HDAC2 complex represses and Sp1/HIF‐1/p300 complex activates ncx1 gene transcription, in brain ischemia and in ischemic brain preconditioning, by epigenetic mechanism. J Neurosci. 2015;35:7332–7348. doi: 10.1523/JNEUROSCI.2174-14.2015 - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

