Bioinformatics analysis and experimental validation of key genes associated with lumbar disc degeneration and biomechanics
- PMID: 38463775
- PMCID: PMC10920361
- DOI: 10.1016/j.heliyon.2024.e27016
Bioinformatics analysis and experimental validation of key genes associated with lumbar disc degeneration and biomechanics
Abstract
Background: Lumbar disc degeneration (LDD) is an important pathological basis for the development of degenerative diseases of the lumbar spine. Most clinical patients have low back pain as their main symptom. The deterioration of the biomechanical environment is an important cause of LDD. Although there is a large amount of basic research on LDD, there are fewer reports that correlate biomechanical mechanisms with basic research. Our research aims to identify 304 key genes involved in LDD due to biomechanical deterioration, using a bioinformatics approach. We focus on SMAD3, CAV1, SMAD7, TGFB1 as hub genes, and screen for 30 potential target drugs, offering novel insights into LDD pathology and treatment options.
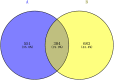
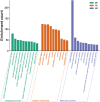
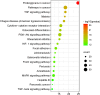
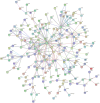
Methods: The Gene Cards, GenCLip3, OMIM and Drugbank databases were explored to obtain genes associated with biomechanics and LDD, followed by making veen plots to obtain both co-expressed genes. GO enrichment analysis and KEGG pathway analysis of the co-expressed genes were obtained using the DAVID online platform and visualised via a free online website. Protein interaction networks (PPI) were obtained through the STRING platform and visualised through Cytoscape 3.9.0. These genes were predicted for downstream interaction networks using the STITCH platform. Then, the GSE56081 dataset was used to validate the key genes. RT-PCR was used to detect mRNA expression of core genes in the degenerated nucleus pulposus (NP) samples and western bolt was used for protein expression. Lastly, the obtained hub genes were searched in the drug database (DGIdb) to find relevant drug candidates.
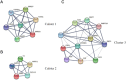
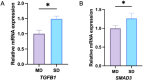
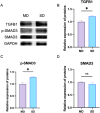
Results: From the perspective of biomechanics-induced LDD, we obtained a total of 304 genes, the GO functional enrichment and KEGG pathway enrichment analysis showed that the functions of these genes are mostly related to inflammation and apoptosis. The PPI network was constructed and four Hub genes were obtained through the plug-in of Cytoscape software, namely SMAD3, CAV1, SMAD7 and TGFB1. The analysis of key genes revealed that biomechanical involvement in LDD may be related to the TGF-β signaling pathway. Validation of the GSE56081 dataset revealed that SMAD3 and TGFB1 were highly expressed in degenerating NP samples. RT-PCR results showed that the mRNA expression of SMAD3 and TGFB1 was significantly increased in the severe degeneration group; Western blot results also showed that the protein expression of TGFB1 and P-SMAD3 was significantly increased. In addition, we identified 30 potential drugs.
Conclusion: This study presented a new approach to investigate the correlation between biomechanical mechanisms and LDD. The deterioration of the biomechanical environment may cause LDD through the TGF-β signaling pathway. TGFB1 and SMAD3 are important core targets. The important genes, pathways and drugs obtained in this study provided a new basis and direction for the study, diagnosis and treatment of LDD.
Keywords: Bioinformatics; Biomechanics; Genes; Lumbar disc degeneration; Targeted drugs.
© 2024 The Authors.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures




Similar articles
-
Comprehensive analysis of lumbar disc degeneration and autophagy-related candidate genes, pathways, and targeting drugs.J Orthop Surg Res. 2021 Apr 13;16(1):252. doi: 10.1186/s13018-021-02417-2. J Orthop Surg Res. 2021. PMID: 33849578 Free PMC article.
-
Identification of common pathway and hub genes in the degeneration of both annulus fibrosus and nucleus pulposus in intervertebral disc.J Orthop Surg (Hong Kong). 2023 Jan-Apr;31(1):10225536231167705. doi: 10.1177/10225536231167705. J Orthop Surg (Hong Kong). 2023. PMID: 36972403
-
Identification of key potential targets for TNF-α/TNFR1-related intervertebral disc degeneration by bioinformatics analysis.Connect Tissue Res. 2021 Sep;62(5):531-541. doi: 10.1080/03008207.2020.1797709. Epub 2020 Aug 26. Connect Tissue Res. 2021. PMID: 32686499
-
Identification of Core Genes and Screening of Potential Targets in Intervertebral Disc Degeneration Using Integrated Bioinformatics Analysis.Front Genet. 2022 May 30;13:864100. doi: 10.3389/fgene.2022.864100. eCollection 2022. Front Genet. 2022. PMID: 35711934 Free PMC article.
-
Research trend of MRI application for lumbar disc degeneration with low back pain: a bibliometric analysis.Front Neurol. 2024 Apr 17;15:1360091. doi: 10.3389/fneur.2024.1360091. eCollection 2024. Front Neurol. 2024. PMID: 38694782 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Miscellaneous

