This is a preprint.
Linewidth-related bias in modelled concentration estimates from GABA-edited 1H-MRS
- PMID: 38464094
- PMCID: PMC10925149
- DOI: 10.1101/2024.02.27.582249
Linewidth-related bias in modelled concentration estimates from GABA-edited 1H-MRS
Abstract
J-difference-edited MRS is widely used to study GABA in the human brain. Editing for low-concentration target molecules (such as GABA) typically exhibits lower signal-to-noise ratio (SNR) than conventional non-edited MRS, varying with acquisition region, volume and duration. Moreover, spectral lineshape may be influenced by age-, pathology-, or brain-region-specific effects of metabolite T2, or by task-related blood-oxygen level dependent (BOLD) changes in functional MRS contexts. Differences in both SNR and lineshape may have systematic effects on concentration estimates derived from spectral modelling. The present study characterises the impact of lineshape and SNR on GABA+ estimates from different modelling algorithms: FSL-MRS, Gannet, LCModel, Osprey, spant and Tarquin. Publicly available multi-site GABA-edited data (222 healthy subjects from 20 sites; conventional MEGA-PRESS editing; TE = 68 ms) were pre-processed with a standardised pipeline, then filtered to apply controlled levels of Lorentzian and Gaussian linebroadening and SNR reduction. Increased Lorentzian linewidth was associated with a 2-5% decrease in GABA+ estimates per Hz, observed consistently (albeit to varying degrees) across datasets and most algorithms. Weaker, often opposing effects were observed for Gaussian linebroadening. Variations are likely caused by differing baseline parametrization and lineshape constraints between models. Effects of linewidth on other metabolites (e.g., Glx and tCr) varied, suggesting that a linewidth confound may persist after scaling to an internal reference. These findings indicate a potentially significant confound for studies where linewidth may differ systematically between groups or experimental conditions, e.g. due to T2 differences between brain regions, age, or pathology, or varying T2* due to BOLD-related changes. We conclude that linewidth effects need to be rigorously considered during experimental design and data processing, for example by incorporating linewidth into statistical analysis of modelling outcomes or development of appropriate lineshape matching algorithms.
Keywords: GABA; MEGA-PRESS; MRS; functional spectroscopy; linewidth; quantification; spectral editing.
Conflict of interest statement
Declaration of Interest The authors declare no conflicting interests.
Figures
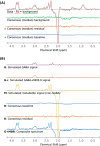






References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
