This is a preprint.
Metformin Enhances Antibody-Mediated Recognition of HIV-Infected CD4+ T-Cells by Decreasing Viral Release
- PMID: 38464135
- PMCID: PMC10925111
- DOI: 10.1101/2024.02.15.580166
Metformin Enhances Antibody-Mediated Recognition of HIV-Infected CD4+ T-Cells by Decreasing Viral Release
Abstract
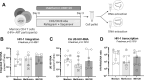
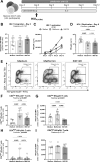
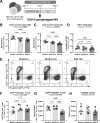
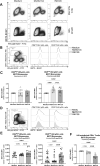
The mechanistic target of rapamycin (mTOR) positively regulates multiple steps of the HIV-1 replication cycle. We previously reported that a 12-weeks supplementation of antiretroviral therapy (ART) with metformin, an indirect mTOR inhibitor used in type-2 diabetes treatment, reduced mTOR activation and HIV transcription in colon-infiltrating CD4+ T-cells, together with systemic inflammation in nondiabetic people with HIV-1 (PWH). Herein, we investigated the antiviral mechanisms of metformin. In a viral outgrowth assay performed with CD4+ T-cells from ART-treated PWH, and upon infection in vitro with replication-competent and VSV-G-pseudotyped HIV-1, metformin decreased virion release, but increased the frequency of productively infected CD4lowHIV-p24+ T-cells. These observations coincided with increased BST2/Tetherin (HIV release inhibitor) and Bcl-2 (pro-survival factor) expression, and improved recognition of productively infected T-cells by HIV-1 Envelope antibodies. Thus, metformin exerts pleiotropic effects on post-transcription/translation steps of the HIV-1 replication cycle and may be used to accelerate viral reservoir decay in ART-treated PWH.
Conflict of interest statement
CONFLICT OF INTEREST The authors declare no competing interests.
Figures



Similar articles
-
Metformin facilitates viral reservoir reactivation and their recognition by anti-HIV-1 envelope antibodies.iScience. 2024 Aug 5;27(9):110670. doi: 10.1016/j.isci.2024.110670. eCollection 2024 Sep 20. iScience. 2024. PMID: 39252967 Free PMC article.
-
LILAC pilot study: Effects of metformin on mTOR activation and HIV reservoir persistence during antiretroviral therapy.EBioMedicine. 2021 Mar;65:103270. doi: 10.1016/j.ebiom.2021.103270. Epub 2021 Mar 1. EBioMedicine. 2021. PMID: 33662832 Free PMC article. Clinical Trial.
-
Choice of antiretroviral therapy differentially impacts survival of HIV-infected CD4 T cells.Mol Cell Ther. 2014 Jan 3;2:1. doi: 10.1186/2052-8426-2-1. eCollection 2014. Mol Cell Ther. 2014. PMID: 26057236 Free PMC article.
-
Reservoirs for HIV-1: mechanisms for viral persistence in the presence of antiviral immune responses and antiretroviral therapy.Annu Rev Immunol. 2000;18:665-708. doi: 10.1146/annurev.immunol.18.1.665. Annu Rev Immunol. 2000. PMID: 10837072 Review.
-
Episomal HIV-1 DNA and its relationship to other markers of HIV-1 persistence.Retrovirology. 2018 Jan 30;15(1):15. doi: 10.1186/s12977-018-0398-1. Retrovirology. 2018. PMID: 29378611 Free PMC article. Review.
References
-
- Samaras K., Wand H., Law M., Emery S., Cooper D., and Carr A. (2007). Prevalence of metabolic syndrome in HIV-infected patients receiving highly active antiretroviral therapy using International Diabetes Foundation and Adult Treatment Panel III criteria: associations with insulin resistance, disturbed body fat compartmentalization, elevated C-reactive protein, and [corrected] hypoadiponectinemia. Diabetes Care 30, 113–119. 10.2337/dc06-1075. - DOI - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
