This is a preprint.
Magnesium Transporter MgtA revealed as a Dimeric P-type ATPase
- PMID: 38464158
- PMCID: PMC10925321
- DOI: 10.1101/2024.02.28.582502
Magnesium Transporter MgtA revealed as a Dimeric P-type ATPase
Update in
-
P-type ATPase magnesium transporter MgtA acts as a dimer.Nat Struct Mol Biol. 2025 Jun 23. doi: 10.1038/s41594-025-01593-7. Online ahead of print. Nat Struct Mol Biol. 2025. PMID: 40550995
Abstract
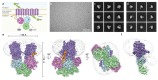
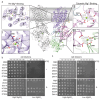
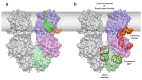
Magnesium (Mg2+) uptake systems are present in all domains of life given the vital role of this ion. Bacteria acquire Mg2+ via conserved Mg2+ channels and transporters. The transporters are required for growth when Mg2+ is limiting or during bacterial pathogenesis, but, despite their significance, there are no known structures for these transporters. Here we report the first structure of the Mg2+ transporter MgtA solved by single particle cryo-electron microscopy (cryo-EM). Using mild membrane extraction, we obtained high resolution structures of both a homodimeric form (2.9 Å), the first for a P-type ATPase, and a monomeric form (3.6 Å). Each monomer unit of MgtA displays a structural architecture that is similar to other P-type ATPases with a transmembrane domain and two soluble domains. The dimer interface consists of contacts between residues in adjacent soluble nucleotide binding and phosphotransfer regions of the haloacid dehalogenase (HAD) domain. We suggest oligomerization is a conserved structural feature of the diverse family of P-type ATPase transporters. The ATP binding site and conformational dynamics upon nucleotide binding to MgtA were characterized using a combination of cryo-EM, molecular dynamics simulations, hydrogen-deuterium exchange mass spectrometry, and mutagenesis. Our structure also revealed a Mg2+ ion in the transmembrane segments, which, when combined with sequence conservation and mutagenesis studies, allowed us to propose a model for Mg2+ transport across the lipid bilayer. Finally, our work revealed the N-terminal domain structure and cytoplasmic Mg2+ binding sites, which have implications for related P-type ATPases defective in human disease.
Keywords: P-type ATPase; cryo-EM; dimer; ion translocation; magnesium; membrane protein; oligomer; transport.
Conflict of interest statement
Competing interests The authors declare no competing interests.
Figures


Similar articles
-
P-type ATPase magnesium transporter MgtA acts as a dimer.Nat Struct Mol Biol. 2025 Jun 23. doi: 10.1038/s41594-025-01593-7. Online ahead of print. Nat Struct Mol Biol. 2025. PMID: 40550995
-
Cholinesterase inhibitors for vascular dementia and other vascular cognitive impairments: a network meta-analysis.Cochrane Database Syst Rev. 2021 Feb 22;2(2):CD013306. doi: 10.1002/14651858.CD013306.pub2. Cochrane Database Syst Rev. 2021. PMID: 33704781 Free PMC article.
-
Intravenous magnesium sulphate and sotalol for prevention of atrial fibrillation after coronary artery bypass surgery: a systematic review and economic evaluation.Health Technol Assess. 2008 Jun;12(28):iii-iv, ix-95. doi: 10.3310/hta12280. Health Technol Assess. 2008. PMID: 18547499
-
Corticosteroids for the treatment of Duchenne muscular dystrophy.Cochrane Database Syst Rev. 2016 May 5;2016(5):CD003725. doi: 10.1002/14651858.CD003725.pub4. Cochrane Database Syst Rev. 2016. PMID: 27149418 Free PMC article.
-
Sertindole for schizophrenia.Cochrane Database Syst Rev. 2005 Jul 20;2005(3):CD001715. doi: 10.1002/14651858.CD001715.pub2. Cochrane Database Syst Rev. 2005. PMID: 16034864 Free PMC article.
References
-
- Jin F., Huang Y. & Hattori M. Recent advances in the structural biology of Mg2+ channels and transporters. J. Mol. Biol. 434, 167729 (2022). - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
