Lipoarabinomannan mediates localized cell wall integrity during division in mycobacteria
- PMID: 38467648
- PMCID: PMC10928101
- DOI: 10.1038/s41467-024-46565-5
Lipoarabinomannan mediates localized cell wall integrity during division in mycobacteria
Abstract
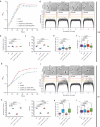
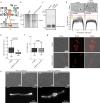
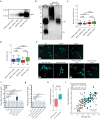
The growth and division of mycobacteria, which include clinically relevant pathogens, deviate from that of canonical bacterial models. Despite their Gram-positive ancestry, mycobacteria synthesize and elongate a diderm envelope asymmetrically from the poles, with the old pole elongating more robustly than the new pole. The phosphatidylinositol-anchored lipoglycans lipomannan (LM) and lipoarabinomannan (LAM) are cell envelope components critical for host-pathogen interactions, but their physiological functions in mycobacteria remained elusive. In this work, using biosynthetic mutants of these lipoglycans, we examine their roles in maintaining cell envelope integrity in Mycobacterium smegmatis and Mycobacterium tuberculosis. We find that mutants defective in producing mature LAM fail to maintain rod cell shape specifically at the new pole and para-septal regions whereas a mutant that produces a larger LAM becomes multi-septated. Therefore, LAM plays critical and distinct roles at subcellular locations associated with division in mycobacteria, including maintenance of local cell wall integrity and septal placement.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Update of
-
Lipoarabinomannan regulates septation in Mycobacterium smegmatis.bioRxiv [Preprint]. 2023 Mar 26:2023.03.26.534150. doi: 10.1101/2023.03.26.534150. bioRxiv. 2023. Update in: Nat Commun. 2024 Mar 11;15(1):2191. doi: 10.1038/s41467-024-46565-5. PMID: 36993273 Free PMC article. Updated. Preprint.
Similar articles
-
Lipoarabinomannan regulates septation in Mycobacterium smegmatis.bioRxiv [Preprint]. 2023 Mar 26:2023.03.26.534150. doi: 10.1101/2023.03.26.534150. bioRxiv. 2023. Update in: Nat Commun. 2024 Mar 11;15(1):2191. doi: 10.1038/s41467-024-46565-5. PMID: 36993273 Free PMC article. Updated. Preprint.
-
Critical roles for lipomannan and lipoarabinomannan in cell wall integrity of mycobacteria and pathogenesis of tuberculosis.mBio. 2013 Feb 19;4(1):e00472-12. doi: 10.1128/mBio.00472-12. mBio. 2013. PMID: 23422411 Free PMC article.
-
Role of LmeA, a Mycobacterial Periplasmic Protein, in Maintaining the Mannosyltransferase MptA and Its Product Lipomannan under Stress.mSphere. 2020 Nov 4;5(6):e01039-20. doi: 10.1128/mSphere.01039-20. mSphere. 2020. PMID: 33148829 Free PMC article.
-
Mycobacterial lipoarabinomannan and related lipoglycans: from biogenesis to modulation of the immune response.Mol Microbiol. 2004 Jul;53(2):391-403. doi: 10.1111/j.1365-2958.2004.04183.x. Mol Microbiol. 2004. PMID: 15228522 Review.
-
Lipoarabinomannan, and its related glycolipids, induce divergent and opposing immune responses to Mycobacterium tuberculosis depending on structural diversity and experimental variations.Tuberculosis (Edinb). 2016 Jan;96:120-30. doi: 10.1016/j.tube.2015.09.005. Epub 2015 Oct 28. Tuberculosis (Edinb). 2016. PMID: 26586646 Review.
Cited by
-
The mycobacterial glycoside hydrolase LamH enables capsular arabinomannan release and stimulates growth.Nat Commun. 2024 Jul 9;15(1):5740. doi: 10.1038/s41467-024-50051-3. Nat Commun. 2024. PMID: 38982040 Free PMC article.
-
Exploiting thioether reactivity to label mycobacterial glycans.Proc Natl Acad Sci U S A. 2025 May 13;122(19):e2422185122. doi: 10.1073/pnas.2422185122. Epub 2025 May 5. Proc Natl Acad Sci U S A. 2025. PMID: 40324093
-
Antigen-presenting innate lymphoid cells induced by BCG vaccination promote a respiratory antiviral immune response through the skin‒lung axis.Cell Mol Immunol. 2025 Apr;22(4):390-402. doi: 10.1038/s41423-025-01267-w. Epub 2025 Feb 17. Cell Mol Immunol. 2025. PMID: 39962263
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

