Comprehensive study reveals phenotypic heterogeneity in Klebsiella pneumoniae species complex isolates
- PMID: 38467675
- PMCID: PMC10928225
- DOI: 10.1038/s41598-024-55546-z
Comprehensive study reveals phenotypic heterogeneity in Klebsiella pneumoniae species complex isolates
Abstract
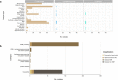
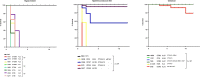
Here, we conducted a comprehensive analysis of 356 Klebsiella pneumoniae species complex (KpSC) isolates that were classified as classical (cl), presumptive hypervirulent (p-hv) and hypermucoviscous-like (hmv-like). Overall, K. pneumoniae (82.3%), K. variicola (2.5%) and K. quasipneumoniae (2.5%) were identified. These isolates comprised 321 cl-KpSC, 7 p-hv-KpSC and 18 hmv-like-KpSC. A large proportion of cl-KpSC isolates were extended-spectrum-β-lactamases (ESBLs)-producers (64.4%) and 3.4% of isolates were colistin-resistant carrying carbapenemase and ESBL genes. All p-hv-KpSC showed an antibiotic susceptible phenotype and hmv-like isolates were found to be ESBL-producers (8/18). Assays for capsule production and capsule-dependent virulence phenotypes and whole-genome sequencing (WGS) were performed in a subset of isolates. Capsule amount differed in all p-hv strains and hmv-like produced higher capsule amounts than cl strains; these variations had important implications in phagocytosis and virulence. Murine sepsis model showed that most cl strains were nonlethal and the hmv-like caused 100% mortality with 3 × 108 CFUs. Unexpectedly, 3/7 (42.9%) of p-hv strains required 108 CFUs to cause 100% mortality (atypical hypervirulent), and 4/7 (57.1%) strains were considered truly hypervirulent (hv). Genomic analyses confirmed the diverse population, including isolates belonging to hv clonal groups (CG) CG23, CG86, CG380 and CG25 (this corresponded to the ST3999 a novel hv clone) and MDR clones such as CG258 and CG147 (ST392) among others. We noted that the hmv-like and hv-ST3999 isolates showed a close phylogenetic relationship with cl-MDR K. pneumoniae. The information collected here is important to understand the evolution of clinically important phenotypes such as hypervirulent and ESBL-producing-hypermucoviscous-like amongst the KpSC in Mexican healthcare settings. Likewise, this study shows that mgrB inactivation is the main mechanism of colistin resistance in K. pneumoniae isolates from Mexico.
Keywords: Capsule; Colistin-resistance; Hypermucoviscosity; Plasmids; Virulence.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
In vivo evolution to hypermucoviscosity and ceftazidime/avibactam resistance in a liver abscess caused by Klebsiella pneumoniae sequence type 512.mSphere. 2024 Sep 25;9(9):e0042324. doi: 10.1128/msphere.00423-24. Epub 2024 Aug 22. mSphere. 2024. PMID: 39171923 Free PMC article.
-
Colistin-resistant Klebsiella pneumoniae species complex: The scenario in Mexico.J Glob Antimicrob Resist. 2025 Jun;43:203-212. doi: 10.1016/j.jgar.2025.04.024. Epub 2025 May 5. J Glob Antimicrob Resist. 2025. PMID: 40334840
-
Multidrug-resistant ESBL-producing Klebsiella pneumoniae complex in Czech hospitals, wastewaters and surface waters.Antimicrob Resist Infect Control. 2024 Nov 26;13(1):141. doi: 10.1186/s13756-024-01496-0. Antimicrob Resist Infect Control. 2024. PMID: 39593189 Free PMC article.
-
Emergence of hypermucoviscous colistin-resistant high-risk convergent Klebsiella pneumoniae ST-2096 clone from Pakistan.Future Microbiol. 2022 Sep;17:989-1000. doi: 10.2217/fmb-2021-0292. Epub 2022 Jul 21. Future Microbiol. 2022. PMID: 35860964
-
Limited Transmission of Klebsiella pneumoniae among Humans, Animals, and the Environment in a Caribbean Island, Guadeloupe (French West Indies).Microbiol Spectr. 2022 Oct 26;10(5):e0124222. doi: 10.1128/spectrum.01242-22. Epub 2022 Sep 12. Microbiol Spectr. 2022. PMID: 36094181 Free PMC article.
Cited by
-
Prevalence and molecular characteristics of colistin-resistant isolates among carbapenem-resistant Klebsiella pneumoniae in Central South China: a multicenter study.Ann Clin Microbiol Antimicrob. 2025 Jan 4;24(1):1. doi: 10.1186/s12941-024-00769-1. Ann Clin Microbiol Antimicrob. 2025. PMID: 39755702 Free PMC article.
-
Complete genome sequences of Klebsiella pneumoniae, Klebsiella quasipneumoniae, and Klebsiella variicola clinical isolates from an epidemiology study.Microbiol Resour Announc. 2025 Mar 11;14(3):e0106024. doi: 10.1128/mra.01060-24. Epub 2025 Feb 12. Microbiol Resour Announc. 2025. PMID: 39936930 Free PMC article.
-
Molecular epidemiology and clinical features of Klebsiella variicola bloodstream infection compared with infection with other Klebsiella pneumoniae species complex strains.Microbiol Spectr. 2025 Jun 3;13(6):e0301724. doi: 10.1128/spectrum.03017-24. Epub 2025 Apr 25. Microbiol Spectr. 2025. PMID: 40277351 Free PMC article.
-
KpSC-ID: a multiplex real-time PCR assay for the simultaneous detection of the Klebsiella pneumoniae species complex and specific identification of Klebsiella pneumoniae, Klebsiella quasipneumoniae and Klebsiella variicola.Microbiology (Reading). 2025 Jul;171(7):001587. doi: 10.1099/mic.0.001587. Microbiology (Reading). 2025. PMID: 40737176 Free PMC article.
-
Pyogenic liver abscess caused by an atypical hypervirulent Klebsiella pneumoniae K1-ST23 in Mexico.IDCases. 2024 May 12;36:e01987. doi: 10.1016/j.idcr.2024.e01987. eCollection 2024. IDCases. 2024. PMID: 38779143 Free PMC article.
References
-
- World Health Organization. WHO publishes list of bacteria for which new antibiotics are urgently needed. https://www.who.int/news/item/27-02-2017-who-publishes-list-of-bacteria-..., 2023 (accessed 28 August 2023).
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

