CTX-M-127 with I176F mutations found in bacteria isolates from Bangladeshi circulating banknotes
- PMID: 38467683
- PMCID: PMC10928162
- DOI: 10.1038/s41598-024-56207-x
CTX-M-127 with I176F mutations found in bacteria isolates from Bangladeshi circulating banknotes
Abstract
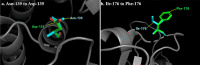
Extended-spectrum beta-lactamase (ESBL)-producing organisms are widely recognized as clinically relevant causes of difficult-to-treat infections. CTX-M has formed a rapidly growing family distributed worldwide among a wide range of clinical bacteria, particularly members of Enterobacteriaceae. Circulating banknotes, exchanged daily among people, pose a potential vehicle for transmitting multidrug resistance. We screened for ESBL-carrying bacteria in the present study and reported CTX-M mutations in Bangladesh's banknotes. We sequenced the genes and performed homology modeling using the Swiss model with CTX-M-15 (4HBT) as a template. Then, we performed molecular docking of mecillinam with the template and the generated model using Autodock 4.2 (Release 4.2.6). After docking, we visually inspected the complexes built using Autodock tools for polar contacts and pi-pi interactions in PyMOL 2.5.4. Our partially sequenced blaCTX-M was related to blaCTX-M-10 and blaCTX-M-15. We observed multiple single-nucleotide substitution mutations, i.e., G613T (silent mutation), A626T (I176F), and A503G (N135D). Homology modeling showed high similarity when the model was superimposed over the template. The orientation of Asn (135) in the template and Asp (135) in the model does not show a significant difference. Likewise, Ile (176) in the template and Phe (176) in the model offer the same orientation. Our generated model could bind to Lys237, Ser240, and Asp135 residues with the lowest binding energy on docking. Our predicted binding of the mecillinam to the mutated D-135 residue in the model indicates contributions and supports previous reports proposing CTX-M-15 to CTX-M-127 mutational conversion on the mecillinum resistance phenotype.
Keywords: Antibiotic resistance; Banknotes; CTX-M-127; CTX-M-15; Extended-spectrum beta-lactamase; Mutation.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



References
-
- Okesola, A.O. & Makanjuola O. Resistance to third-generation Cephalosporins and other antibiotics by Enterobacteriaceae in Western Nigeria. Am. J. Infect. Diseases. 5(1), 17–20 (2009).
-
- Barthelemy M, Peduzzi J, Bernard H, Tancrede C, Labia R. Close amino acid sequence relationship between the new plasmid-mediated extended-spectrum beta-lactamase MEN-1 and chromosomally encoded enzymes of Klebsiella oxytoca. Biochim. Biophys. Acta. 1992;1122(1):15–22. doi: 10.1016/0167-4838(92)90121-S. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

