Identification of potential immunotherapeutic targets and prognostic biomarkers in Graves' disease using weighted gene co-expression network analysis
- PMID: 38468967
- PMCID: PMC10926144
- DOI: 10.1016/j.heliyon.2024.e27175
Identification of potential immunotherapeutic targets and prognostic biomarkers in Graves' disease using weighted gene co-expression network analysis
Abstract
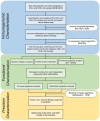
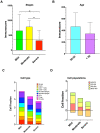
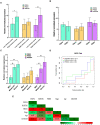
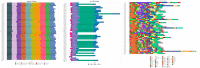
Graves' disease (GD) is an autoimmune disorder characterized by hyperthyroidism resulting from autoantibody-induced stimulation of the thyroid gland. Despite recent advancements in understanding GD's pathogenesis, the molecular processes driving disease progression and treatment response remain poorly understood. In this study, we aimed to identify crucial immunogenic factors associated with GD prognosis and immunotherapeutic response. To achieve this, we implemented a comprehensive screening strategy that combined computational immunogenicity-potential scoring with multi-parametric cluster analysis to assess the immunomodulatory genes in GD-related subtypes involving stromal and immune cells. Utilizing weighted gene co-expression network analysis (WGCNA), we identified co-expressed gene modules linked to cellular senescence and immune infiltration in CD4+ and CD8+ GD samples. Additionally, gene set enrichment analysis enabled the identification of hallmark pathways distinguishing high- and low-immune subtypes. Our WGCNA analysis revealed 21 gene co-expression modules comprising 1,541 genes associated with immune infiltration components in various stages of GD, including T cells, M1 and M2 macrophages, NK cells, and Tregs. These genes primarily participated in T cell proliferation through purinergic signaling pathways, particularly neuroactive ligand-receptor interactions, and DNA binding transcription factor activity. Three genes, namely PRSS1, HCRTR1, and P2RY4, exhibited robustness in GD patients across multiple stages and were involved in immune cell infiltration during the late stage of GD (p < 0.05). Importantly, HCRTR1 and P2RY4 emerged as potential prognostic signatures for predicting overall survival in high-immunocore GD patients (p < 0.05). Overall, our study provides novel insights into the molecular mechanisms driving GD progression and highlights potential key immunogens for further investigation. These findings underscore the significance of immune infiltration-related cellular senescence in GD therapy and present promising targets for the development of new immunotherapeutic strategies.
Keywords: Gene co-expression modules; Graves' disease; Immune infiltration; Immunotherapeutic biomarkers; Molecular mechanisms; WGCNA.
© 2024 Published by Elsevier Ltd.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures



Similar articles
-
T-cell exhaustion-related genes in Graves' disease: a comprehensive genome mapping analysis.Front Endocrinol (Lausanne). 2024 Aug 22;15:1364782. doi: 10.3389/fendo.2024.1364782. eCollection 2024. Front Endocrinol (Lausanne). 2024. PMID: 39239096 Free PMC article.
-
Key gene co-expression modules and functional pathways involved in the pathogenesis of Graves' disease.Mol Cell Endocrinol. 2018 Oct 15;474:252-259. doi: 10.1016/j.mce.2018.03.015. Epub 2018 Mar 31. Mol Cell Endocrinol. 2018. PMID: 29614339
-
A cellular senescence-related classifier based on a tumorigenesis- and immune infiltration-guided strategy can predict prognosis, immunotherapy response, and candidate drugs in hepatocellular carcinoma.Front Immunol. 2022 Nov 15;13:974377. doi: 10.3389/fimmu.2022.974377. eCollection 2022. Front Immunol. 2022. PMID: 36458010 Free PMC article.
-
The Role of the Microbiota in Graves' Disease and Graves' Orbitopathy.Front Cell Infect Microbiol. 2021 Dec 22;11:739707. doi: 10.3389/fcimb.2021.739707. eCollection 2021. Front Cell Infect Microbiol. 2021. PMID: 35004341 Free PMC article. Review.
-
Regulatory B and T cell responses in patients with autoimmune thyroid disease and healthy controls.Dan Med J. 2016 Feb;63(2):B5177. Dan Med J. 2016. PMID: 26836805 Review.
References
-
- Antonelli A., Fallahi P., Elia G., Ragusa F., Paparo S.R., Ruffilli I., et al. Graves' disease: clinical manifestations, immune pathogenesis (cytokines and chemokines) and therapy. Best Pract. Res. Clin. Endocrinol. Metabol. 2020;34(1) doi: 10.1016/j.beem.2020.101388. Epub 2020/02/16. PubMed PMID: 32059832. - DOI - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

