Comparative transcriptomic analysis reveals differences in gene expression and regulatory pathways between nonacral and acral melanoma in Asian individuals
- PMID: 38469735
- PMCID: PMC11484150
- DOI: 10.1111/1346-8138.17187
Comparative transcriptomic analysis reveals differences in gene expression and regulatory pathways between nonacral and acral melanoma in Asian individuals
Abstract
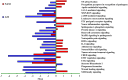
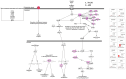
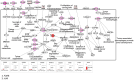
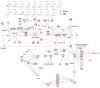
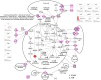
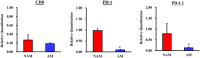
Melanoma predominantly occurs in White individuals, which is associated with factors such as exposure to UV radiation and skin pigmentation. Despite its low incidence, melanoma is the primary cause of skin cancer-related death in Asia, typically in areas with low sun exposure. In our previous whole-exome sequencing study, we identified mutational signature 12 as the most prevalent variant in Asian patients, differing from the common UV-associated mutational signature 7 observed in White individuals. We also observed major differences between acral melanoma (AM) and nonacral melanoma (NAM) in terms of signatures 7, 21, and 22. Notably, few studies have investigated the genomic differences between AM and NAM in Asian individuals. Therefore, in this study, we conducted transcriptomic sequencing to examine the disparities in RNA expression between AM and NAM. Ribosomal RNA depletion was performed to enhance the detection of functionally relevant coding and noncoding transcripts. Ingenuity pathway analysis revealed significant differences in gene expression and regulatory pathways between AM and NAM. The results also indicate that the genes involved in cell cycle signaling or immune modulation and programmed cell death protein 1/programmed cell death 1 ligand 1 signaling were differentially expressed in NAM and AM. In addition, high CDK4 expression and cell cycle variability were observed in AM, with high immunogenicity in NAM. Overall, these findings provide further insights into the pathogenesis of melanoma and serve as a reference for future research on this major malignant disease.
Keywords: Asians; RNA sequencing; melanoma; transcriptome.
© 2024 The Authors. The Journal of Dermatology published by John Wiley & Sons Australia, Ltd on behalf of Japanese Dermatological Association.
Conflict of interest statement
None declared.
Figures


References
-
- Bolick NL, Geller AC. Epidemiology of melanoma. Hematol/Oncol Clin. 2021;35:57–72. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

