A vaccine targeting antigen-presenting cells through CD40 induces protective immunity against Nipah disease
- PMID: 38471503
- PMCID: PMC10983108
- DOI: 10.1016/j.xcrm.2024.101467
A vaccine targeting antigen-presenting cells through CD40 induces protective immunity against Nipah disease
Abstract
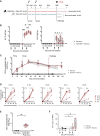
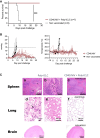
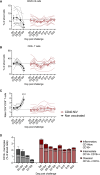
Nipah virus (NiV) has been recently ranked by the World Health Organization as being among the top eight emerging pathogens likely to cause major epidemics, whereas no therapeutics or vaccines have yet been approved. We report a method to deliver immunogenic epitopes from NiV through the targeting of the CD40 receptor of antigen-presenting cells by fusing a selected humanized anti-CD40 monoclonal antibody to the Nipah glycoprotein with conserved NiV fusion and nucleocapsid peptides. In the African green monkey model, CD40.NiV induces specific immunoglobulin A (IgA) and IgG as well as cross-neutralizing responses against circulating NiV strains and Hendra virus and T cell responses. Challenge experiments using a NiV-B strain demonstrate the high protective efficacy of the vaccine, with all vaccinated animals surviving and showing no significant clinical signs or virus replication, suggesting that the CD40.NiV vaccine conferred sterilizing immunity. Overall, results obtained with the CD40.NiV vaccine are highly promising in terms of the breadth and efficacy against NiV.
Keywords: CD40-targeting vaccine; Henipavirus; Nipah virus; cross-neutralizing responses; dendritic cell-based vaccine; epitope mapping; germinal center reaction; neutralizing antibodies; prophylactic vaccine; sterilizing immunity.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures



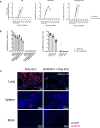

Similar articles
-
Single-dose live-attenuated Nipah virus vaccines confer complete protection by eliciting antibodies directed against surface glycoproteins.Vaccine. 2014 May 7;32(22):2637-44. doi: 10.1016/j.vaccine.2014.02.087. Epub 2014 Mar 12. Vaccine. 2014. PMID: 24631094 Free PMC article.
-
Use of Single-Injection Recombinant Vesicular Stomatitis Virus Vaccine to Protect Nonhuman Primates Against Lethal Nipah Virus Disease.Emerg Infect Dis. 2019 Jun;25(6):1144-1152. doi: 10.3201/eid2506.181620. Emerg Infect Dis. 2019. PMID: 31107231 Free PMC article.
-
Immunological correlates of protection afforded by PHV02 live, attenuated recombinant vesicular stomatitis virus vector vaccine against Nipah virus disease.Front Immunol. 2023 Sep 4;14:1216225. doi: 10.3389/fimmu.2023.1216225. eCollection 2023. Front Immunol. 2023. PMID: 37731485 Free PMC article.
-
Advancements in Nipah virus treatment: Analysis of current progress in vaccines, antivirals, and therapeutics.Immunology. 2024 Feb;171(2):155-169. doi: 10.1111/imm.13695. Epub 2023 Sep 15. Immunology. 2024. PMID: 37712243 Review.
-
Nipah virus: epidemiology, pathology, immunobiology and advances in diagnosis, vaccine designing and control strategies - a comprehensive review.Vet Q. 2019 Dec;39(1):26-55. doi: 10.1080/01652176.2019.1580827. Vet Q. 2019. PMID: 31006350 Free PMC article. Review.
Cited by
-
Development of nebulized inhalation delivery for fusion-inhibitory lipopeptides to protect non-human primates against Nipah-Bangladesh infection.Antiviral Res. 2025 Mar;235:106095. doi: 10.1016/j.antiviral.2025.106095. Epub 2025 Jan 25. Antiviral Res. 2025. PMID: 39870114 Free PMC article.
-
Henipaviruses: epidemiology, ecology, disease, and the development of vaccines and therapeutics.Clin Microbiol Rev. 2025 Mar 13;38(1):e0012823. doi: 10.1128/cmr.00128-23. Epub 2024 Dec 23. Clin Microbiol Rev. 2025. PMID: 39714175 Free PMC article. Review.
-
Risk of Nipah Virus Seroprevalence in Healthcare Workers: A Systematic Review with Meta-Analysis.Viruses. 2025 Jan 9;17(1):81. doi: 10.3390/v17010081. Viruses. 2025. PMID: 39861870 Free PMC article.
-
Emerging and reemerging infectious diseases: global trends and new strategies for their prevention and control.Signal Transduct Target Ther. 2024 Sep 11;9(1):223. doi: 10.1038/s41392-024-01917-x. Signal Transduct Target Ther. 2024. PMID: 39256346 Free PMC article. Review.
-
Targeting Langerhans cells via skin delivery of HIV Envelope enhances the antibody response to vaccination.NPJ Vaccines. 2025 Jul 25;10(1):170. doi: 10.1038/s41541-025-01214-w. NPJ Vaccines. 2025. PMID: 40715153 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

