Whole-Genome Resequencing of Ujimqin Sheep Identifies Genes Associated with Vertebral Number
- PMID: 38473062
- PMCID: PMC10931107
- DOI: 10.3390/ani14050677
Whole-Genome Resequencing of Ujimqin Sheep Identifies Genes Associated with Vertebral Number
Abstract
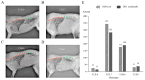
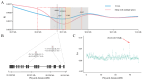
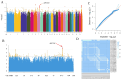
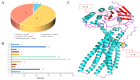
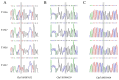
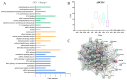
The number of vertebrae is a crucial economic trait that can significantly impact the carcass length and meat production in animals. However, our understanding of the quantitative trait loci (QTLs) and candidate genes associated with the vertebral number in sheep (Ovis aries) remains limited. To identify these candidate genes and QTLs, we collected 73 Ujimqin sheep with increased numbers of vertebrae (T13L7, T14L6, and T14L7) and 23 sheep with normal numbers of vertebrae (T13L6). Through high-throughput genome resequencing, we obtained a total of 24,130,801 effective single-nucleotide polymorphisms (SNPs). By conducting a selective-sweep analysis, we discovered that the most significantly selective region was located on chromosome 7. Within this region, we identified several genes, including VRTN, SYNDIG1L, LTBP2, and ABCD4, known to regulate the spinal development and morphology. Further, a genome-wide association study (GWAS) performed on sheep with increased and normal vertebral numbers confirmed that ABCD4 is a candidate gene for determining the number of vertebrae in sheep. Additionally, the most significant SNP on chromosome 7 was identified as a candidate QTL. Moreover, we detected two missense mutations in the ABCD4 gene; one of these mutations (Chr7: 89393414, C > T) at position 22 leads to the conversion of arginine (Arg) to glutamine (Gln), which is expected to negatively affect the protein's function. Notably, a transcriptome expression profile in mouse embryonic development revealed that ABCD4 is highly expressed during the critical period of vertebral formation (4.5-7.5 days). Our study highlights ABCD4 as a potential major gene influencing the number of vertebrae in Ujimqin sheep, with promising prospects for future genome-assisted breeding improvements in sheep.
Keywords: ABCD4 gene; Ujimqin sheep; vertebral number; whole-genome resequencing.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
Similar articles
-
Rapid Identification of the SNP Mutation in the ABCD4 Gene and Its Association with Multi-Vertebrae Phenotypes in Ujimqin Sheep Using TaqMan-MGB Technology.Animals (Basel). 2025 Aug 5;15(15):2284. doi: 10.3390/ani15152284. Animals (Basel). 2025. PMID: 40805073 Free PMC article.
-
Whole-Genome Resequencing Reveals Loci Associated With Thoracic Vertebrae Number in Sheep.Front Genet. 2019 Jul 18;10:674. doi: 10.3389/fgene.2019.00674. eCollection 2019. Front Genet. 2019. PMID: 31379930 Free PMC article.
-
Understanding molecular mechanisms of vertebral number of variations on Mongolian sheep using candidate genes analysis.Anim Biosci. 2025 Feb;38(2):247-254. doi: 10.5713/ab.24.0212. Epub 2024 Aug 26. Anim Biosci. 2025. PMID: 39210812 Free PMC article.
-
Effects of vertebral number variations on carcass traits and genotyping of Vertnin candidate gene in Kazakh sheep.Asian-Australas J Anim Sci. 2017 Sep;30(9):1234-1238. doi: 10.5713/ajas.16.0959. Epub 2017 Mar 25. Asian-Australas J Anim Sci. 2017. PMID: 28423880 Free PMC article.
-
Review on Genomic Regions and Candidate Genes Associated with Economically Important Production and Reproduction Traits in Sheep (Ovies aries).Animals (Basel). 2019 Dec 23;10(1):33. doi: 10.3390/ani10010033. Animals (Basel). 2019. PMID: 31877963 Free PMC article. Review.
Cited by
-
Potential candidate genes influencing meat production phenotypic traits in sheep: a review.Front Vet Sci. 2025 Jul 16;12:1616533. doi: 10.3389/fvets.2025.1616533. eCollection 2025. Front Vet Sci. 2025. PMID: 40740300 Free PMC article. Review.
-
Rapid Identification of the SNP Mutation in the ABCD4 Gene and Its Association with Multi-Vertebrae Phenotypes in Ujimqin Sheep Using TaqMan-MGB Technology.Animals (Basel). 2025 Aug 5;15(15):2284. doi: 10.3390/ani15152284. Animals (Basel). 2025. PMID: 40805073 Free PMC article.
-
Leveraging Whole-Genome Resequencing to Uncover Genetic Diversity and Promote Conservation Strategies for Ruminants in Asia.Animals (Basel). 2025 Mar 13;15(6):831. doi: 10.3390/ani15060831. Animals (Basel). 2025. PMID: 40150358 Free PMC article. Review.
References
-
- King J.W.B., Roberts R.C. Carcass Length in the Bacon Pig; Its Association with Vertebrae Numbers and Prediction from Radiographs of the Young Pig. Anim. Prod. 1960;2:59–65. doi: 10.1017/S0003356100033493. - DOI
Grants and funding
- 62061034, 62171241/The National Nature Scientific Foundation of China
- 2020GG0055, 2020CG0078, 2019GG241, 219GG348, 2020ZD0007, 2021GG0062/Science and Technology Planning Project of Inner Mongolia
- 2021GG0398/The key technology research program of Inner Mongolia Autonomous Region
- 2019ZD031/The Science and Technology Major Project of Inner Mongolia Autonomous Region of China to the State Key Laboratory of Reproductive Regulation and Breeding of Grassland Livestock
LinkOut - more resources
Full Text Sources
Miscellaneous

