Genome-Wide Transcriptome Profiling Reveals the Mechanisms Underlying Hepatic Metabolism under Different Raising Systems in Yak
- PMID: 38473080
- PMCID: PMC10930694
- DOI: 10.3390/ani14050695
Genome-Wide Transcriptome Profiling Reveals the Mechanisms Underlying Hepatic Metabolism under Different Raising Systems in Yak
Abstract
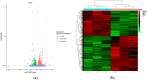
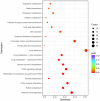
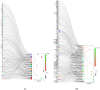
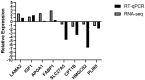
Yak meat is nutritionally superior to beef cattle but has a low fat content and is slow-growing. The liver plays a crucial role in lipid metabolism, and in order to determine whether different feeding modes affect lipid metabolism in yaks and how it is regulated, we employed RNA sequencing (RNA-seq) technology to analyze the genome-wide differential gene expression in the liver of yaks maintained under different raising systems. A total of 1663 differentially expressed genes (DEGs) were identified (|log2FC| ≥ 0 and p-value ≤ 0.05), including 698 down-regulated and 965 up-regulated genes. According to gene ontology (GO) and KEGG enrichment analyses, these DEGs were significantly enriched in 13 GO terms and 26 pathways (p < 0.05). Some DEGs were enriched in fatty acid degradation, PPAR, PI3K-Akt, and ECM receptor pathways, which are associated with lipid metabolism. A total of 16 genes are well known to be related to lipid metabolism (e.g., APOA1, FABP1, EHHADH, FADS2, SLC27A5, ACADM, CPT1B, ACOX2, HMGCS2, PLIN5, ACAA1, IGF1, FGFR4, ALDH9A1, ECHS1, LAMA2). A total of 11 of the above genes were significantly enriched in the PPAR signaling pathway. The reliability of the transcriptomic data was verified using qRT-PCR. Our findings provide new insights into the mechanisms regulating yak meat quality. It shows that fattening improves the expression of genes that regulate lipid deposition in yaks and enhances meat quality. This finding will contribute to a better understanding of the various factors that determine yak meat quality and help develop strategies to improve yield and quality.
Keywords: RNA-seq; fattening; lipid metabolism; liver; yak meat.
Conflict of interest statement
The authors declare no conflicts of interest. The author who is not in the same organization as the other authors, and the organization he belongs to do not have a conflict of interest with this experiment.
Figures


Similar articles
-
RNA-Seq Analysis Identifies Differentially Expressed Genes Insubcutaneous Adipose Tissuein Qaidamford Cattle, Cattle-Yak, and Angus Cattle.Animals (Basel). 2019 Dec 3;9(12):1077. doi: 10.3390/ani9121077. Animals (Basel). 2019. PMID: 31816988 Free PMC article.
-
Identification of differentially expressed genes at different post-natal development stages of longissimus dorsi muscle in Tianzhu white yak.Gene. 2022 May 20;823:146356. doi: 10.1016/j.gene.2022.146356. Epub 2022 Feb 25. Gene. 2022. PMID: 35227854
-
RNA-Seq Analysis Identifies Differentially Expressed Genes in the Longissimus dorsi of Wagyu and Chinese Red Steppe Cattle.Int J Mol Sci. 2022 Dec 26;24(1):387. doi: 10.3390/ijms24010387. Int J Mol Sci. 2022. PMID: 36613828 Free PMC article.
-
The Effect of the Feeding System on Fat Deposition in Yak Subcutaneous Fat.Int J Mol Sci. 2023 Apr 17;24(8):7381. doi: 10.3390/ijms24087381. Int J Mol Sci. 2023. PMID: 37108542 Free PMC article.
-
Long-stranded non-coding RNAs temporal-specific expression profiles reveal longissimus dorsi muscle development and intramuscular fat deposition in Tianzhu white yak.J Anim Sci. 2023 Jan 3;101:skad394. doi: 10.1093/jas/skad394. J Anim Sci. 2023. PMID: 38029315 Free PMC article.
Cited by
-
Insights into Adaption and Growth Evolution: Genome-Wide Copy Number Variation Analysis in Chinese Hainan Yellow Cattle Using Whole-Genome Re-Sequencing Data.Int J Mol Sci. 2024 Nov 6;25(22):11919. doi: 10.3390/ijms252211919. Int J Mol Sci. 2024. PMID: 39595990 Free PMC article.
-
Integrated Transcriptomic Analysis of Liver and Muscle Tissues Reveals Candidate Genes and Pathways Regulating Intramuscular Fat Deposition in Beef Cattle.Animals (Basel). 2025 Apr 30;15(9):1306. doi: 10.3390/ani15091306. Animals (Basel). 2025. PMID: 40362121 Free PMC article.
References
-
- Guo X., Long R., Kreuzer M., Ding L., Shang Z., Zhang Y., Yang Y., Cui G. Importance of functional ingredients in yak milk-derived food on health of Tibetan nomads living under high-altitude stress: A review. Crit. Rev. Food Sci. Nutr. 2014;54:292–302. doi: 10.1080/10408398.2011.584134. - DOI - PubMed
-
- Li Y., Zong W., Zhao S., Qie M., Yang X., Zhao Y. Nutrition and edible characteristics, origin traceability and authenticity identification of yak meat and milk: A review. Trends Food Sci. Technol. 2023;139:104133. doi: 10.1016/j.tifs.2023.104133. - DOI
Grants and funding
- 2022YFD1302101/State Key R & D program
- QYXTZX-NY2022, QYXTZX-RKZ2022/Regional innovation project of Tibet Autonomous Region
- 20JR5RA580/Gansu basic research innovation group project
- MATS-Beef Cattle System, CARS-37/Modern beef yak industry technology system
- 25-LIHPS-01/Yak resources and breeding innovation project of Chinese Academy of Agricultural Sciences
LinkOut - more resources
Full Text Sources
Miscellaneous

