Exploring the Molecular Tapestry: Organ-Specific Peptide and Protein Ultrafiltrates and Their Role in Therapeutics
- PMID: 38474110
- PMCID: PMC10931619
- DOI: 10.3390/ijms25052863
Exploring the Molecular Tapestry: Organ-Specific Peptide and Protein Ultrafiltrates and Their Role in Therapeutics
Abstract
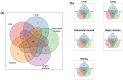
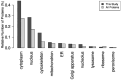
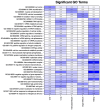
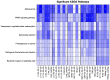
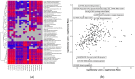
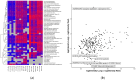
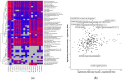
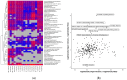
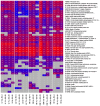
This study aims to characterize the proteome composition of organ-derived protein extracts from rabbits. Protein isolation was performed using soft homogenization and size exclusion via ultrafiltration. The proteome analysis of the ultrafiltrates was conducted using gel electrophoresis, and the mass spectrometry data were subjected to gene ontology analysis. Proteomic profiling revealed comprehensive protein profiles associated with RNA regulation, fatty acid binding, inflammatory response, oxidative stress, and metabolism. Additionally, our results demonstrate the presence of abundant small proteins, as observed in the mass spectrometry datasets. Small proteins and peptides are crucial in transcription modulation and various biological processes. The protein networks identified in the ultrafiltrates have the potential to enhance and complement biological therapeutic interventions. Data are available via ProteomeXchange with identifier PXD050039.
Keywords: organ specificity; proteomics; small proteins; ultrafiltration.
Conflict of interest statement
Authors affiliated with the European Wellness Academy and European Wellness International are not involved in formal analysis and investigation. The authors have no other relevant affiliations or financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript apart from those disclosed.
Figures



Similar articles
-
Human follicular fluid proteomic and peptidomic composition quantitative studies by SWATH-MS methodology. Applicability of high pH RP-HPLC fractionation.J Proteomics. 2019 Jan 16;191:131-142. doi: 10.1016/j.jprot.2018.03.010. Epub 2018 Mar 10. J Proteomics. 2019. PMID: 29530678
-
Proteomic profiling of heat acclimation in cerebrospinal fluid of rabbit.J Proteomics. 2016 Jul 20;144:113-22. doi: 10.1016/j.jprot.2016.04.028. Epub 2016 May 18. J Proteomics. 2016. PMID: 27208788
-
Characterization of the Low-Molecular-Weight Human Plasma Peptidome.Methods Mol Biol. 2017;1619:63-79. doi: 10.1007/978-1-4939-7057-5_6. Methods Mol Biol. 2017. PMID: 28674878
-
Emerging Affinity-Based Proteomic Technologies for Large-Scale Plasma Profiling in Cardiovascular Disease.Circulation. 2017 Apr 25;135(17):1651-1664. doi: 10.1161/CIRCULATIONAHA.116.025446. Circulation. 2017. PMID: 28438806 Free PMC article. Review.
-
Sample preparation by in-gel digestion for mass spectrometry-based proteomics.Anal Bioanal Chem. 2007 Oct;389(4):991-1002. doi: 10.1007/s00216-007-1451-4. Epub 2007 Jul 17. Anal Bioanal Chem. 2007. PMID: 17639354 Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

