DPPA2/4 Promote the Pluripotency and Proliferation of Bovine Extended Pluripotent Stem Cells by Upregulating the PI3K/AKT/GSK3β/β-Catenin Signaling Pathway
- PMID: 38474345
- PMCID: PMC10930381
- DOI: 10.3390/cells13050382
DPPA2/4 Promote the Pluripotency and Proliferation of Bovine Extended Pluripotent Stem Cells by Upregulating the PI3K/AKT/GSK3β/β-Catenin Signaling Pathway
Abstract
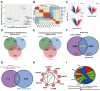
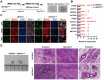
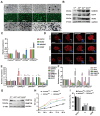
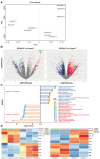
Developmental pluripotency-associated 2 (DPPA2) and DPPA4 are crucial transcription factors involved in maintaining pluripotency in humans and mice. However, the role of DPPA2/4 in bovine extended pluripotent stem cells (bEPSCs) has not been investigated. In this study, a subset of bEPSC-related differentially expressed genes (DEGs), including DPPA2 and DPPA4, was identified based on multiomics data (ATAC-seq and RNA-seq). Subsequent investigations revealed that double overexpression of DPPA2/4 facilitates the reprogramming of bovine fetal fibroblasts (BFFs) into bEPSCs, whereas knockout of DPPA2/4 in BFFs leads to inefficient reprogramming. DPPA2/4 overexpression and knockdown experiments revealed that the pluripotency and proliferation capability of bEPSCs were maintained by promoting the transition from the G1 phase to the S phase of the cell cycle. By activating the PI3K/AKT/GSK3β/β-catenin pathway in bEPSCs, DPPA2/4 can increase the nuclear accumulation of β-catenin, which further upregulates lymphoid enhancer binding factor 1 (LEF1) transcription factor activity. Moreover, DPPA2/4 can also regulate the expression of LEF1 by directly binding to its promoter region. Overall, our results demonstrate that DPPA2/4 promote the reprogramming of BFFs into bEPSCs while also maintaining the pluripotency and proliferation capability of bEPSCs by regulating the PI3K/AKT/GSK3β/β-catenin pathway and subsequently activating LEF1. These findings expand our understanding of the gene regulatory network involved in bEPSC pluripotency.
Keywords: DPPA2; DPPA4; LEF1; PI3K/AKT pathway; bEPSCs; pluripotency; proliferation; reprogramming.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures



Similar articles
-
PBX homeobox 1 enhances hair follicle mesenchymal stem cell proliferation and reprogramming through activation of the AKT/glycogen synthase kinase signaling pathway and suppression of apoptosis.Stem Cell Res Ther. 2019 Aug 23;10(1):268. doi: 10.1186/s13287-019-1382-y. Stem Cell Res Ther. 2019. PMID: 31443676 Free PMC article.
-
Dppa2 and Dppa4 directly regulate the Dux-driven zygotic transcriptional program.Genes Dev. 2019 Feb 1;33(3-4):194-208. doi: 10.1101/gad.321174.118. Epub 2019 Jan 28. Genes Dev. 2019. PMID: 30692203 Free PMC article.
-
Identification of DPPA4 and DPPA2 as a novel family of pluripotency-related oncogenes.Stem Cells. 2013 Nov;31(11):2330-42. doi: 10.1002/stem.1526. Stem Cells. 2013. PMID: 23963736 Free PMC article.
-
DPPA2, DPPA4, and other DPPA factor epigenomic functions in cell fate and cancer.Stem Cell Reports. 2021 Dec 14;16(12):2844-2851. doi: 10.1016/j.stemcr.2021.10.008. Epub 2021 Nov 11. Stem Cell Reports. 2021. PMID: 34767751 Free PMC article. Review.
-
Brain-derived Neurotrophic Factor Promotes Growth of Neurons and Neural Stem Cells Possibly by Triggering the Phosphoinositide 3-Kinase/ AKT/Glycogen Synthase Kinase-3β/β-catenin Pathway.CNS Neurol Disord Drug Targets. 2017;16(7):828-836. doi: 10.2174/1871527316666170518170422. CNS Neurol Disord Drug Targets. 2017. PMID: 28524001
Cited by
-
The timing of maternal protein degradation during mammalian preimplantation development is species-specific.Reproduction. 2025 Aug 14;170(3):e250007. doi: 10.1530/REP-25-0007. Print 2025 Sep 1. Reproduction. 2025. PMID: 40812375 Free PMC article.
-
IGFBP7 is a key component of the senescence-associated secretory phenotype (SASP) that induces senescence in healthy cells by modulating the insulin, IGF, and activin A pathways.Cell Commun Signal. 2024 Nov 12;22(1):540. doi: 10.1186/s12964-024-01921-2. Cell Commun Signal. 2024. PMID: 39533382 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
- 32160172/National Natural Science Foundation of China
- 2020BS03003/Natural Science Foundation of Inner Mongolia Autonomous Region
- 2020ZD10/the Major Projects of Natural Science Foundation of Inner Mongolia Autonomous Region
- 2019ZD031/the Science and Technology Major Project of the Inner Mongolia Autonomous Region of China to the State Key Laboratory of Reproductive Regulation
- 2021ZD0048/the Science and Technology Major Project of the Inner Mongolia Autonomous Region of China to the State Key Laboratory of Reproductive Regulation
LinkOut - more resources
Full Text Sources

