Predicting lncRNA-protein interactions through deep learning framework employing multiple features and random forest algorithm
- PMID: 38475723
- PMCID: PMC10929084
- DOI: 10.1186/s12859-024-05727-4
Predicting lncRNA-protein interactions through deep learning framework employing multiple features and random forest algorithm
Abstract
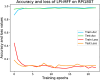
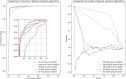
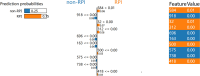
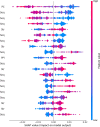
RNA-protein interaction (RPI) is crucial to the life processes of diverse organisms. Various researchers have identified RPI through long-term and high-cost biological experiments. Although numerous machine learning and deep learning-based methods for predicting RPI currently exist, their robustness and generalizability have significant room for improvement. This study proposes LPI-MFF, an RPI prediction model based on multi-source information fusion, to address these issues. The LPI-MFF employed protein-protein interactions features, sequence features, secondary structure features, and physical and chemical properties as the information sources with the corresponding coding scheme, followed by the random forest algorithm for feature screening. Finally, all information was combined and a classification method based on convolutional neural networks is used. The experimental results of fivefold cross-validation demonstrated that the accuracy of LPI-MFF on RPI1807 and NPInter was 97.60% and 97.67%, respectively. In addition, the accuracy rate on the independent test set RPI1168 was 84.9%, and the accuracy rate on the Mus musculus dataset was 90.91%. Accordingly, LPI-MFF demonstrated greater robustness and generalization than other prevalent RPI prediction methods.
Keywords: Features fusion; LncRNA–protein interactions; Multiple features; Random forest algorithm.
© 2024. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


Similar articles
-
RPI-EDLCN: An Ensemble Deep Learning Framework Based on Capsule Network for ncRNA-Protein Interaction Prediction.J Chem Inf Model. 2024 Apr 8;64(7):2221-2235. doi: 10.1021/acs.jcim.3c00377. Epub 2023 May 9. J Chem Inf Model. 2024. PMID: 37158609
-
RPITER: A Hierarchical Deep Learning Framework for ncRNA⁻Protein Interaction Prediction.Int J Mol Sci. 2019 Mar 1;20(5):1070. doi: 10.3390/ijms20051070. Int J Mol Sci. 2019. PMID: 30832218 Free PMC article.
-
LPI-CSFFR: Combining serial fusion with feature reuse for predicting LncRNA-protein interactions.Comput Biol Chem. 2022 Aug;99:107718. doi: 10.1016/j.compbiolchem.2022.107718. Epub 2022 Jun 27. Comput Biol Chem. 2022. PMID: 35785626
-
LMI-DForest: A deep forest model towards the prediction of lncRNA-miRNA interactions.Comput Biol Chem. 2020 Dec;89:107406. doi: 10.1016/j.compbiolchem.2020.107406. Epub 2020 Oct 20. Comput Biol Chem. 2020. PMID: 33120126 Review.
-
LncFinder: an integrated platform for long non-coding RNA identification utilizing sequence intrinsic composition, structural information and physicochemical property.Brief Bioinform. 2019 Nov 27;20(6):2009-2027. doi: 10.1093/bib/bby065. Brief Bioinform. 2019. PMID: 30084867 Free PMC article. Review.
Cited by
-
LncRNA-Protein Interactions: A Key to Deciphering LncRNA Mechanisms.Biomolecules. 2025 Jun 17;15(6):881. doi: 10.3390/biom15060881. Biomolecules. 2025. PMID: 40563521 Free PMC article. Review.
References
-
- Raza A, Uddin J, Almuhaimeed A, Akbar S, Zou Q, Ahmad A. AIPs-SnTCN: Predicting anti-inflammatory peptides using fasttext and transformer encoder-based hybrid word embedding with self-normalized temporal convolutional networks. J Chem Inf Model. 2023;63(21):6537–6554. doi: 10.1021/acs.jcim.3c01563. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

