Transcriptomic and proteomic profiles of fetal versus adult mesenchymal stromal cells and mesenchymal stromal cell-derived extracellular vesicles
- PMID: 38475970
- PMCID: PMC10935839
- DOI: 10.1186/s13287-024-03683-7
Transcriptomic and proteomic profiles of fetal versus adult mesenchymal stromal cells and mesenchymal stromal cell-derived extracellular vesicles
Abstract
Background: Mesenchymal stem/stromal cells (MSCs) can regenerate tissues through engraftment and differentiation but also via paracrine signalling via extracellular vesicles (EVs). Fetal-derived MSCs (fMSCs) have been shown, both in vitro and in animal studies, to be more efficient than adult MSC (aMSCs) in generating bone and muscle but the underlying reason for this difference has not yet been clearly elucidated. In this study, we aimed to systematically investigate the differences between fetal and adult MSCs and MSC-derived EVs at the phenotypic, RNA, and protein levels.
Methods: We carried out a detailed and comparative characterization of culture-expanded fetal liver derived MSCs (fMSCs) and adult bone marrow derived MSCs (aMSCs) phenotypically, and the MSCs and MSC-derived EVs were analysed using transcriptomics and proteomics approaches with RNA Sequencing and Mass Spectrometry.
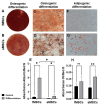
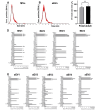
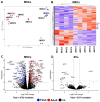
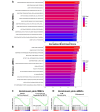
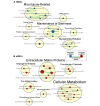
Results: Fetal MSCs were smaller, exhibited increased proliferation and colony-forming capacity, delayed onset of senescence, and demonstrated superior osteoblast differentiation capability compared to their adult counterparts. Gene Ontology analysis revealed that fMSCs displayed upregulated gene sets such as "Positive regulation of stem cell populations", "Maintenance of stemness" and "Muscle cell development/contraction/Myogenesis" in comparison to aMSCs. Conversely, aMSCs displayed upregulated gene sets such as "Complement cascade", "Adipogenesis", "Extracellular matrix glycoproteins" and "Cellular metabolism", and on the protein level, "Epithelial cell differentiation" pathways. Signalling entropy analysis suggested that fMSCs exhibit higher signalling promiscuity and hence, higher potency than aMSCs. Gene ontology comparisons revealed that fetal MSC-derived EVs (fEVs) were enriched for "Collagen fibril organization", "Protein folding", and "Response to transforming growth factor beta" compared to adult MSC-derived EVs (aEVs), whereas no significant difference in protein expression in aEVs compared to fEVs could be detected.
Conclusions: This study provides detailed and systematic insight into the differences between fMSCs and aMSCs, and MSC-derived EVs. The key finding across phenotypic, transcriptomic and proteomic levels is that fMSCs exhibit higher potency than aMSCs, meaning they are in a more undifferentiated state. Additionally, fMSCs and fMSC-derived EVs may possess greater bone forming capacity compared to aMSCs. Therefore, using fMSCs may lead to better treatment efficacy, especially in musculoskeletal diseases.
Keywords: Biosignature; Extracellular vesicles; Fetal mesenchymal stem cells; Mesenchymal stromal cells; Proteomic; Transcriptomic.
© 2024. The Author(s).
Conflict of interest statement
CG and LWJ are co-founders and co-owners of BOOST Pharma ApS. AG and SEA are consultants for and have equity interest in Evox Therapeutics. KLB is consultant for Swedish StromaBio AB. DWH, EBG, FA, MAT, MOG, MP, NS and YKL declare that they have no competing interests.
Figures



References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

