Comparative evolutionary analyses of peste des petits ruminants virus genetic lineages
- PMID: 38476867
- PMCID: PMC10930206
- DOI: 10.1093/ve/veae012
Comparative evolutionary analyses of peste des petits ruminants virus genetic lineages
Abstract
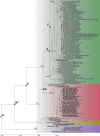
Peste des petits ruminants virus (PPRV) causes a highly infectious disease affecting mainly goats and sheep in large parts of Africa, Asia, and the Middle East and has an important impact on the global economy and food security. Full genome sequencing of PPRV strains has proved to be critical to increasing our understanding of PPR epidemiology and to inform the ongoing global efforts for its eradication. However, the number of full PPRV genomes published is still limited and with a heavy bias towards recent samples and genetic Lineage IV (LIV), which is only one of the four existing PPRV lineages. Here, we generated genome sequences for twenty-five recent (2010-6) and seven historical (1972-99) PPRV samples, focusing mainly on Lineage II (LII) in West Africa. This provided the first opportunity to compare the evolutionary pressures and history between the globally dominant PPRV genetic LIV and LII, which is endemic in West Africa. Phylogenomic analysis showed that the relationship between PPRV LII strains was complex and supported the extensive transboundary circulation of the virus within West Africa. In contrast, LIV sequences were clearly separated per region, with strains from West and Central Africa branched as a sister clade to all other LIV sequences, suggesting that this lineage also has an African origin. Estimates of the time to the most recent common ancestor place the divergence of modern LII and LIV strains in the 1960s-80s, suggesting that this period was particularly important for the diversification and spread of PPRV globally. Phylogenetic relationships among historical samples from LI, LII, and LIII and with more recent samples point towards a high genetic diversity for all these lineages in Africa until the 1970s-80s and possible bottleneck events shaping PPRV's evolution during this period. Molecular evolution analyses show that strains belonging to LII and LIV have evolved under different selection pressures. Differences in codon usage and adaptative selection pressures were observed in all viral genes between the two lineages. Our results confirm that comparative genomic analyses can provide new insights into PPRV's evolutionary history and molecular epidemiology. However, PPRV genome sequencing efforts must be ramped up to increase the resolution of such studies for their use in the development of efficient PPR control and surveillance strategies.
Keywords: endemic; epizootic; evolution; morbillivirus; phylogenetics; small ruminants.
© The Author(s) 2024. Published by Oxford University Press.
Conflict of interest statement
None declared.
Figures


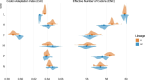

References
-
- Banyard A. C. et al. (2010) ‘Global Distribution of Peste Des Petits Ruminants Virus and Prospects for Improved Diagnosis and Control’, Journal of General Virology, 91: 2885–97. - PubMed
-
- Baron M. D. et al. (2016) ‘Peste Des Petits Ruminants Virus’, in Kielian, M., Maramorosch, K. and Mettenleiter, T. C. (eds) Advances in Virus Research Volume 95, pp. 1–42. Cambridge/Massachusetts/USA: Academic Press. - PubMed
LinkOut - more resources
Full Text Sources

