Hospital antimicrobial stewardship: profiling the oral microbiome after exposure to COVID-19 and antibiotics
- PMID: 38476940
- PMCID: PMC10927822
- DOI: 10.3389/fmicb.2024.1346762
Hospital antimicrobial stewardship: profiling the oral microbiome after exposure to COVID-19 and antibiotics
Abstract
Introduction: During the COVID-19 Delta variant surge, the CLAIRE cross-sectional study sampled saliva from 120 hospitalized patients, 116 of whom had a positive COVID-19 PCR test. Patients received antibiotics upon admission due to possible secondary bacterial infections, with patients at risk of sepsis receiving broad-spectrum antibiotics (BSA).
Methods: The saliva samples were analyzed with shotgun DNA metagenomics and respiratory RNA virome sequencing. Medical records for the period of hospitalization were obtained for all patients. Once hospitalization outcomes were known, patients were classified based on their COVID-19 disease severity and the antibiotics they received.
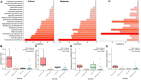
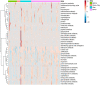
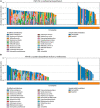
Results: Our study reveals that BSA regimens differentially impacted the human salivary microbiome and disease progression. 12 patients died and all of them received BSA. Significant associations were found between the composition of the COVID-19 saliva microbiome and BSA use, between SARS-CoV-2 genome coverage and severity of disease. We also found significant associations between the non-bacterial microbiome and severity of disease, with Candida albicans detected most frequently in critical patients. For patients who did not receive BSA before saliva sampling, our study suggests Staphylococcus aureus as a potential risk factor for sepsis.
Discussion: Our results indicate that the course of the infection may be explained by both monitoring antibiotic treatment and profiling a patient's salivary microbiome, establishing a compelling link between microbiome and the specific antibiotic type and timing of treatment. This approach can aid with emergency room triage and inpatient management but also requires a better understanding of and access to narrow-spectrum agents that target pathogenic bacteria.
Keywords: COVID-19; Candida albicans (C. albicans); Staphylococcus aureus (S. aureus); broad-spectrum antibiotics; hospital antimicrobial stewardship; saliva microbiome; sepsis.
Copyright © 2024 Buendia, Fernandez, Raley, Rahnavard, Crandall and Castro.
Conflict of interest statement
PB and KF were employed by Lifetime Omics. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The authors declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures



References
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

