Amphiregulin Switches Progenitor Cell Fate for Lineage Commitment During Gastric Mucosal Regeneration
- PMID: 38492892
- PMCID: PMC11260537
- DOI: 10.1053/j.gastro.2024.03.009
Amphiregulin Switches Progenitor Cell Fate for Lineage Commitment During Gastric Mucosal Regeneration
Abstract
Background & aims: Isthmic progenitors, tissue-specific stem cells in the stomach corpus, maintain mucosal homeostasis by balancing between proliferation and differentiation to gastric epithelial lineages. The progenitor cells rapidly adopt an active state in response to mucosal injury. However, it remains unclear how the isthmic progenitor cell niche is controlled during the regeneration of damaged epithelium.
Methods: We recapitulated tissue recovery process after acute mucosal injury in the mouse stomach. Bromodeoxyuridine incorporation was used to trace newly generated cells during the injury and recovery phases. To define the epithelial lineage commitment process during recovery, we performed single-cell RNA-sequencing on epithelial cells from the mouse stomachs. We validated the effects of amphiregulin (AREG) on mucosal recovery, using recombinant AREG treatment or AREG-deficient mice.
Results: We determined that an epidermal growth factor receptor ligand, AREG, can control progenitor cell lineage commitment. Based on the identification of lineage-committed subpopulations in the corpus epithelium through single-cell RNA-sequencing and bromodeoxyuridine incorporation, we showed that isthmic progenitors mainly transition into short-lived surface cell lineages but are less frequently committed to long-lived parietal cell lineages in homeostasis. However, mucosal regeneration after damage directs the lineage commitment of isthmic progenitors towards parietal cell lineages. During recovery, AREG treatment promoted repopulation with parietal cells, while suppressing surface cell commitment of progenitors. In contrast, transforming growth factor-α did not alter parietal cell regeneration, but did induce expansion of surface cell populations. AREG deficiency impairs parietal cell regeneration but increases surface cell commitment.
Conclusions: These data demonstrate that different epidermal growth factor receptor ligands can distinctly regulate isthmic progenitor-driven mucosal regeneration and lineage commitment.
Keywords: Amphiregulin; Epidermal Growth Factor Receptor; Isthmic Progenitor Cell; Lineage Commitment; Parietal Cell Regeneration.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Figures
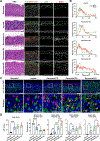






References
-
- Karam SM. Dynamics of epithelial cells in the corpus of the mouse stomach. IV. Bidirectional migration of parietal cells ending in their gradual degeneration and loss. Anat.Rec 1993;236:314–332. - PubMed
-
- Karam SM, Leblond CP. Dynamics of epithelial cells in the corpus of the mouse stomach. I. Identification of proliferative cell types and pinpointing of the stem cells. Anat.Rec 1993;236:259–279. - PubMed
-
- Karam SM, Leblond CP. Dynamics of epithelial cells in the corpus of the mouse stomach. II. Outward migration of pit cells. Anat.Rec 1993;236:280–296. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

