This is a preprint.
Self-organization of embryonic stem cells into a reproducible embryo model through epigenome editing
- PMID: 38496557
- PMCID: PMC10942404
- DOI: 10.1101/2024.03.05.583597
Self-organization of embryonic stem cells into a reproducible embryo model through epigenome editing
Update in
-
Self-organization of mouse embryonic stem cells into reproducible pre-gastrulation embryo models via CRISPRa programming.Cell Stem Cell. 2025 Jun 5;32(6):895-913.e8. doi: 10.1016/j.stem.2025.02.015. Epub 2025 Mar 20. Cell Stem Cell. 2025. PMID: 40118066
Abstract
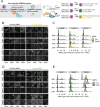
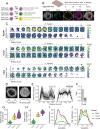
Embryonic stem cells (ESCs) can self-organize in vitro into developmental patterns with spatial organization and molecular similarity to that of early embryonic stages. This self-organization of ESCs requires transmission of signaling cues, via addition of small molecule chemicals or recombinant proteins, to induce distinct embryonic cellular fates and subsequent assembly into structures that can mimic aspects of early embryonic development. During natural embryonic development, different embryonic cell types co-develop together, where each cell type expresses specific fate-inducing transcription factors through activation of non-coding regulatory elements and interactions with neighboring cells. However, previous studies have not fully explored the possibility of engineering endogenous regulatory elements to shape self-organization of ESCs into spatially-ordered embryo models. Here, we hypothesized that cell-intrinsic activation of a minimum number of such endogenous regulatory elements is sufficient to self-organize ESCs into early embryonic models. Our results show that CRISPR-based activation (CRISPRa) of only two endogenous regulatory elements in the genome of pluripotent stem cells is sufficient to generate embryonic patterns that show spatial and molecular resemblance to that of pre-gastrulation mouse embryonic development. Quantitative single-cell live fluorescent imaging showed that the emergence of spatially-ordered embryonic patterns happens through the intrinsic induction of cell fate that leads to an orchestrated collective cellular motion. Based on these results, we propose a straightforward approach to efficiently form 3D embryo models through intrinsic CRISPRa-based epigenome editing and independent of external signaling cues. CRISPRa-Programmed Embryo Models (CPEMs) show highly consistent composition of major embryonic cell types that are spatially-organized, with nearly 80% of the structures forming an embryonic cavity. Single cell transcriptomics confirmed the presence of main embryonic cell types in CPEMs with transcriptional similarity to pre-gastrulation mouse embryos and revealed novel signaling communication links between different embryonic cell types. Our findings offer a programmable embryo model and demonstrate that minimum intrinsic epigenome editing is sufficient to self-organize ESCs into highly consistent pre-gastrulation embryo models.
Conflict of interest statement
Conflict of Interest G.K and R.A are co-founder of Neurosetta, which is focused on commercializing the RosetteArray platform.
Figures



References
-
- Liu X, Tan JP, Schröder J, Aberkane A, Ouyang JF, Mohenska M, et al. Modelling human blastocysts by reprogramming fibroblasts into iBlastoids. Nature. 2021;591: 627–632. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
